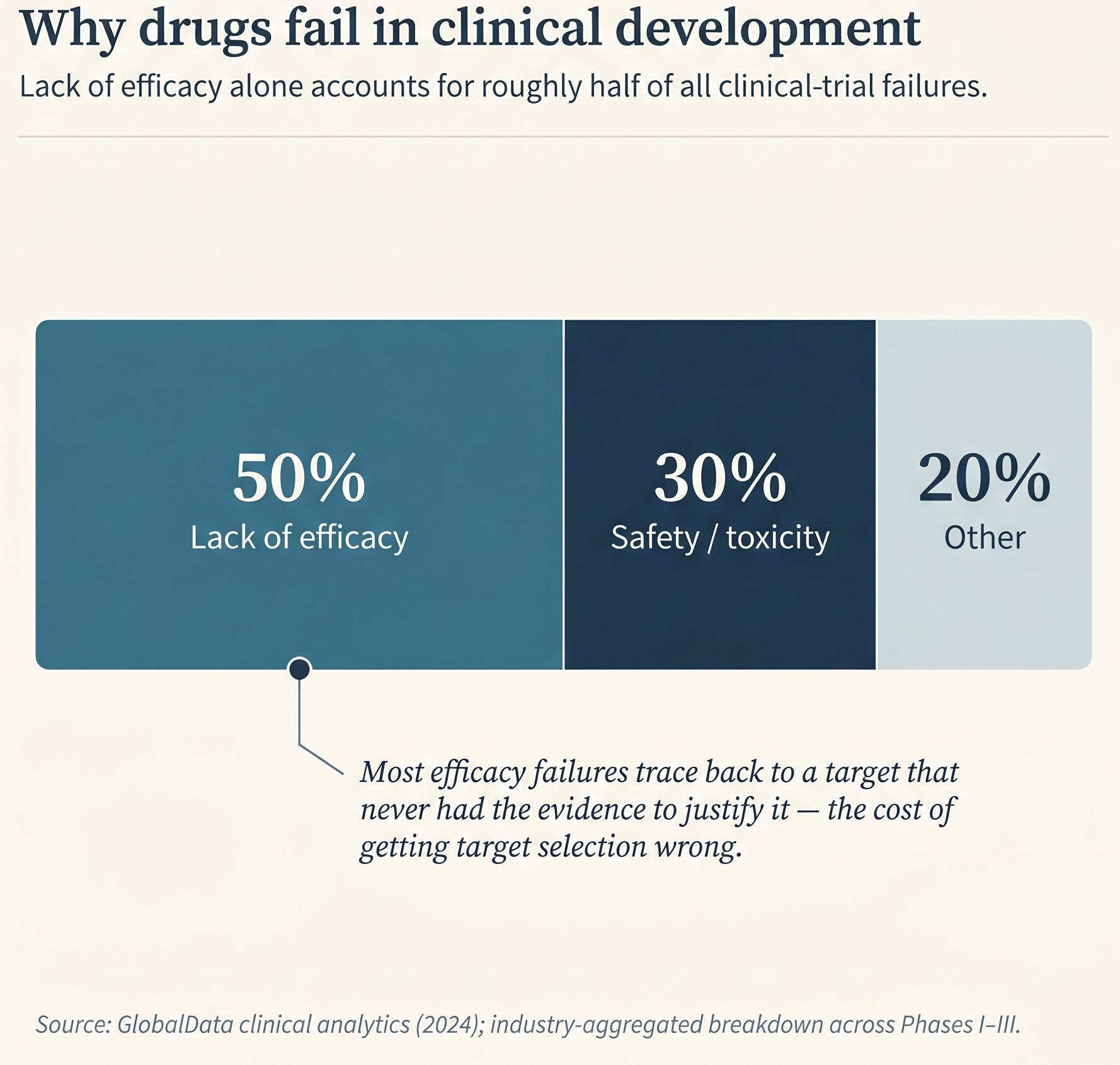
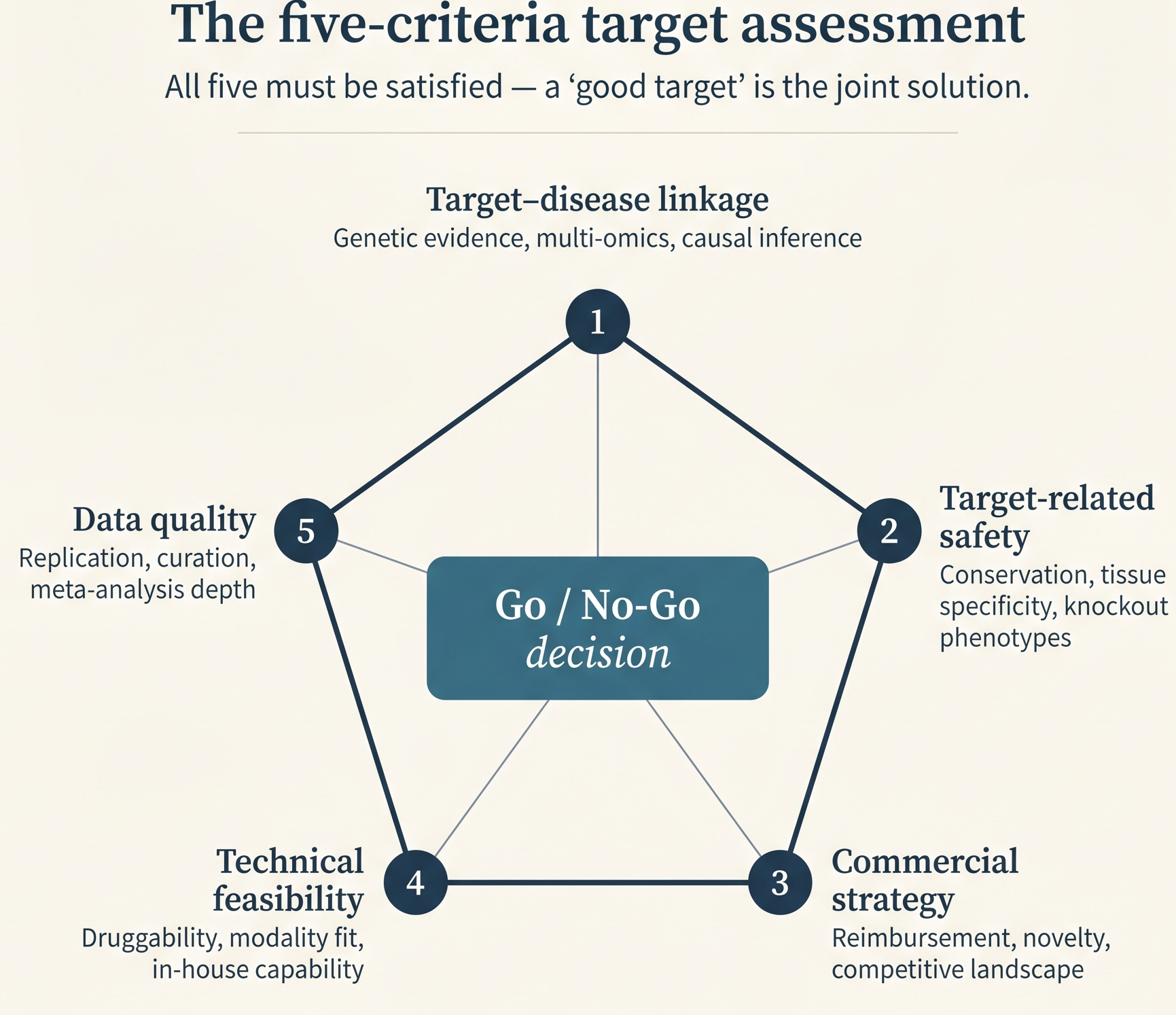
Concept Translation: “Undruggable” is an artifact of the tool - it means the current inhibitor toolkit (small molecules that fit deep pockets) can’t act on this protein. It does not mean the protein is intrinsically resistant to intervention. The ML analogy: a feature you cannot perturb with your current intervention API. Once the API expands (PROTACs, ASOs, ADCs), the same protein moves from undruggable to druggable without any biological change.
Why druggability assessment matters in 2026
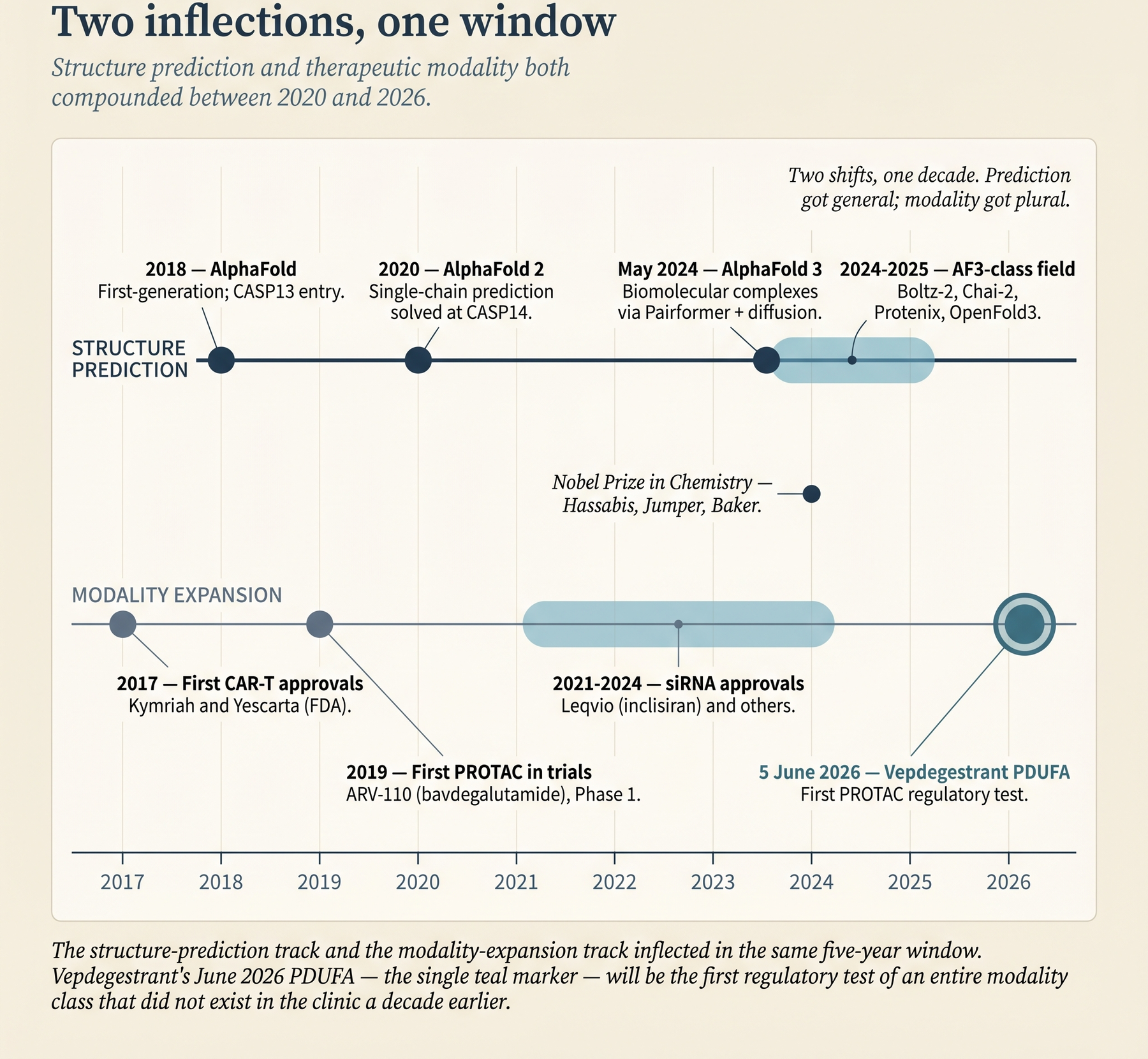
Two shifts have moved the frontier of druggability at roughly the same time, and together they change the analysis.
The first is structural. AlphaFold 2 turned single-chain protein structure prediction from a research problem into routine infrastructure in 2020. AlphaFold 3 in May 2024 extended that capability to biomolecular complexes - proteins with small-molecule ligands, with DNA and RNA, with ions, and with post-translational modifications, all in a single diffusion-based model (Abramson et al., Nature 630:493–500, 2024). The AlphaFold 2 work earned Demis Hassabis and John Jumper a share of the 2024 Nobel Prize in Chemistry, with David Baker taking the other half for computational protein design. AF3 now sits inside a broader field that includes Boltz-1 and Boltz-2 (MIT Jameel Clinic), Chai-1 and Chai-2 (Chai Discovery), Protenix (ByteDance), HelixFold-3 (Baidu), and OpenFold3. Predicted structures are now a default input to virtual screening - the computational ranking of large compound libraries against a target - rather than a special case.
The second shift is modality. Antisense oligonucleotides (ASOs), small interfering RNAs (siRNAs), targeted protein degraders (PROTACs and molecular glues), antibody-drug conjugates (ADCs), and cyclic peptides all have approved products in the market or late-stage assets in development. Each of these expands the “druggable” envelope along a different axis. An RNA-binding ASO can modulate a protein that has no structured binding pocket at all. A PROTAC can take down a scaffolding protein - one that organizes other proteins rather than catalyzing a reaction - even when there is no catalytic activity to inhibit. An ADC can deliver a cytotoxic payload selectively to tumor cells whose surface antigen (a protein displayed on the cell surface) is otherwise non-actionable.
A 2015-era druggability analysis might ask does this target have a small-molecule binding pocket, and is its structure known? A 2026 analysis has to ask that question across at least five modality classes. It also has to answer the harder follow-up: if several modalities are tractable, which one fits this target’s biology, safety profile, and commercial context best?
What “druggable” actually means
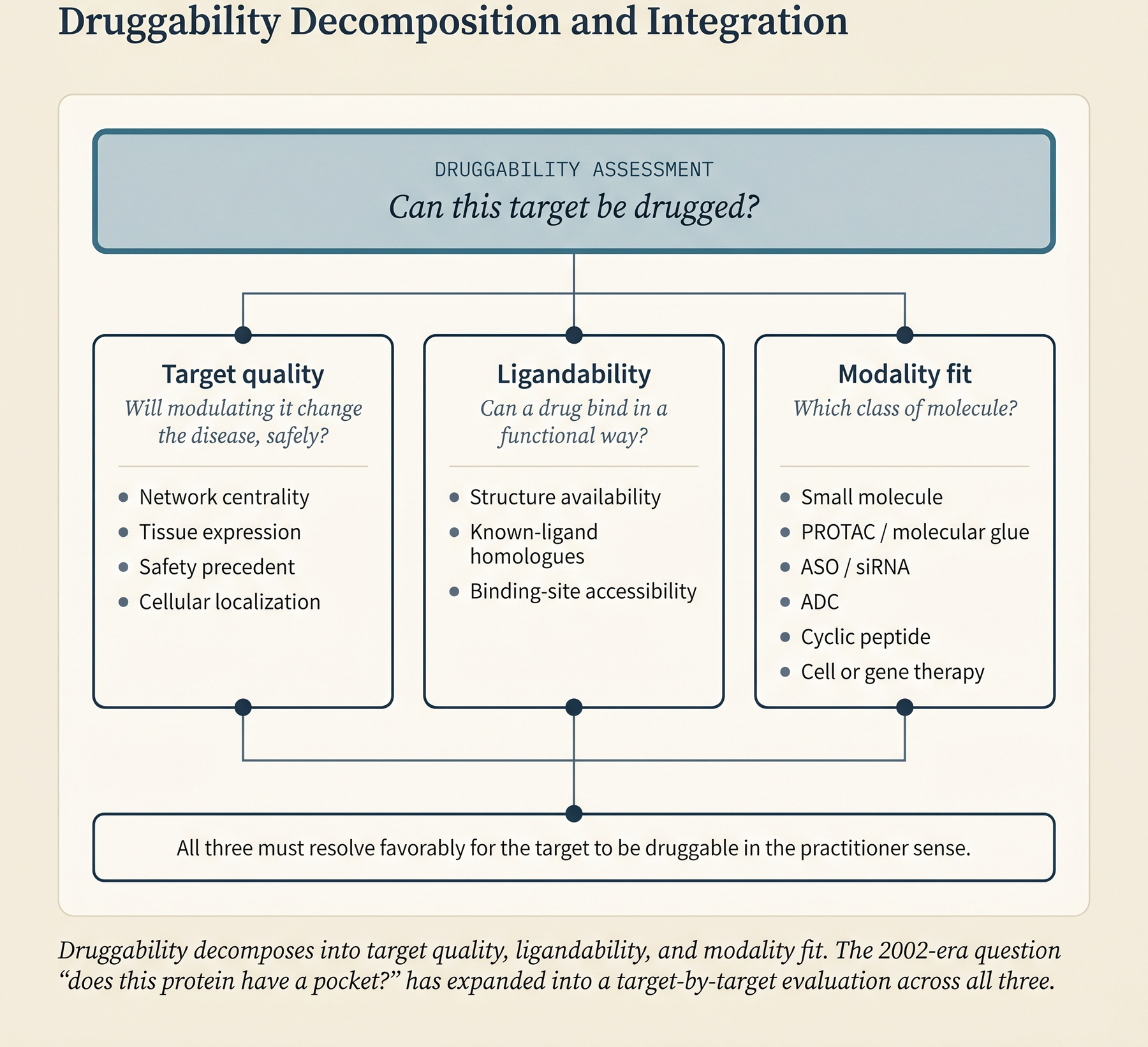
The vocabulary needs care because the field uses these terms loosely.
Druggability refers to the ability of a protein, peptide, or nucleic acid to be modulated by a drug. The 2002 analysis that anchored the field estimated that about 3,000 of the ~20,000 human protein-coding genes - roughly 15% - are druggable in the small-molecule sense, and that only around 27% of those druggable genes actually have an approved drug. Small-molecule drugs target roughly 2–5% of the human genome in total. That figure marks the reach of current chemistry.
Druggability decomposes into two more specific questions.
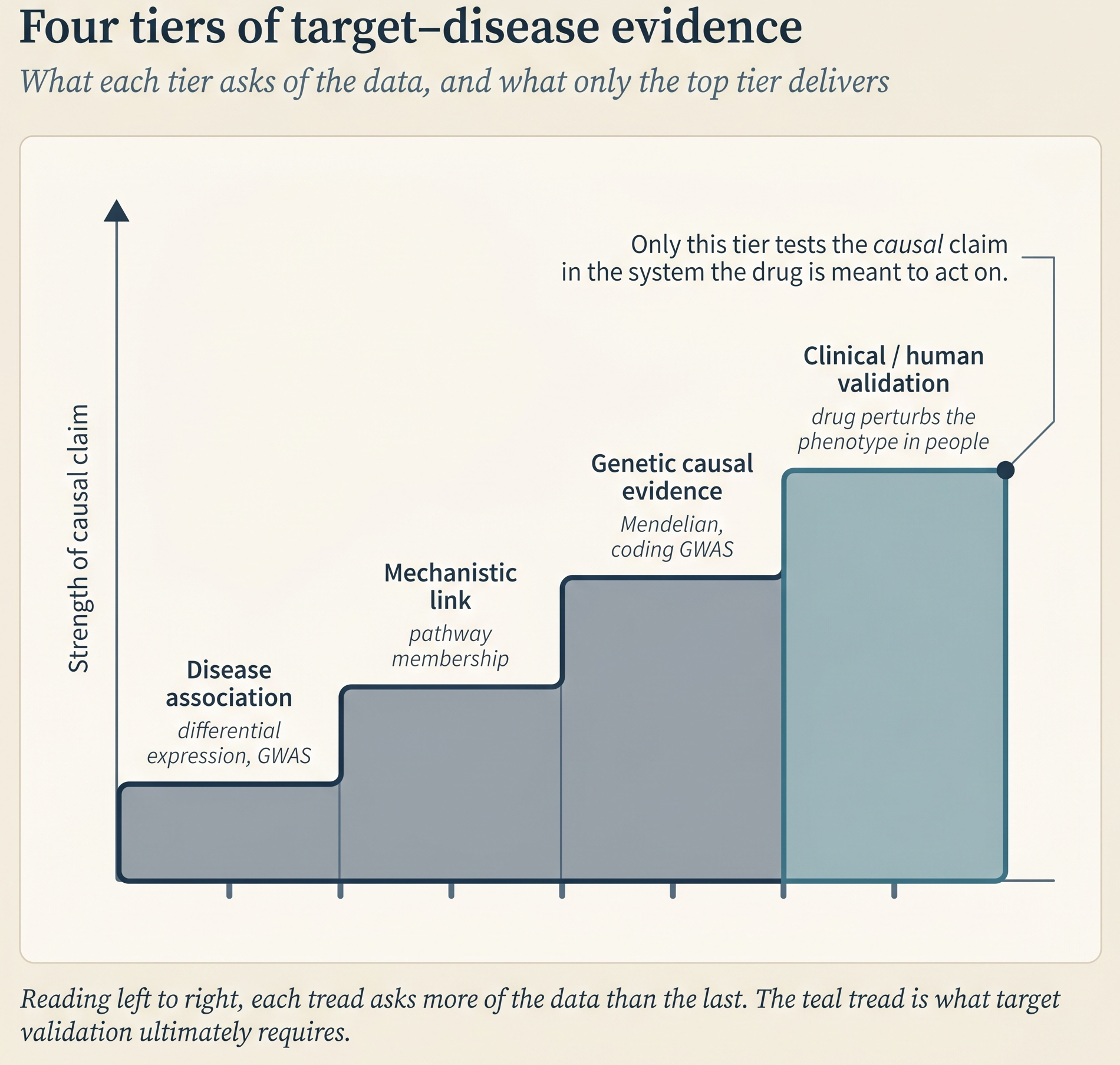
Target quality asks whether modulating the target will actually change the disease, and whether doing so is safe. The relevant variables are the target’s role in essential cellular processes, its network centrality (how many pathways and interactions depend on it), its tissue-expression pattern (a target essential for heart-muscle function is a bad target, almost regardless of its affinity profile), its approval-adjacent precedent (whether closely related targets or modalities have already cleared regulatory review), and its cellular localization (an intracellular target requires permeability that a cell-surface target does not). Target quality is what [target–disease evidence] → How to tell a drug target matters: evidence frameworks for target–disease linkage (C1) gives you the machinery to evaluate.
Ligandability asks whether a drug molecule can bind the target in a way that modulates it. Ligandability depends on three things: the availability of the target’s structure (experimental or predicted), the presence of closely related proteins with known ligands (if a close cousin of your target has been drugged, the odds are better), and the accessibility of a binding site (a small-molecule pocket for small molecules, a regulatory region for nucleic-acid-targeted drugs, or a surface-exposed patch for antibodies).
Concept Translation: Target quality and ligandability are independent axes. A target can be highly ligandable - it has a deep, well-defined pocket, a structure, and a binder - while being a terrible drug target if hitting it doesn’t change disease or causes serious toxicity. Many of the 80–90% of proteins called “undruggable” in older analyses are really not yet ligandable by small molecules; their target quality may be excellent. New modalities are mostly ligandability-extenders, not target-quality-extenders.
Classical small-molecule druggability tools such as DoGSiteScorer (Volkamer et al., 2012), fpocket (Le Guilloux et al., 2009), and SiteMap (Halgren, 2009) turn that third question into a quantitative pocket score. They compute geometric and chemical descriptors of candidate cavities. These remain widely cited workhorses. Newer ML-based approaches (PockDrug, PocketMiner) have emerged and are worth tracking, but the original three are still common reference points in 2026 ligandability analyses. Structure prediction enters the picture at both stages of ligandability: it provides a structure for targets that have never been crystallized, and it increasingly provides one with a candidate ligand already docked - computationally placed into a proposed binding pose.
The classical druggability landscape by protein family
Before structure prediction started moving the boundary, the druggable genome was shaped largely by which protein families had accessible binding sites and tractable chemistry. A handful of families dominate approved drugs.
G protein-coupled receptors (GPCRs) are the single most targeted class. They sit in the cell membrane and relay outside signals inward. Their extracellular ligand-binding domains can be reached by small molecules, peptide-like molecules, antibodies, and allosteric modulators - drugs that bind away from the main ligand site and tune receptor activity.
Ion channels provide a good contrast. The sequenced human genome identified more than 400 putative ion channels, though only a fraction have been cloned and functionally characterized. Their widespread tissue distribution and the physiological consequences of opening or closing them make them both compelling and treacherous targets.
Kinases are the third-most-targeted group. Kinases transfer phosphate groups from ATP to substrate proteins, and the ATP-binding pocket is the canonical small-molecule site. Five inhibitor strategies recur across the literature: ATP mimics (the default), covalent inhibitors (which form a bond with a cysteine in or near the pocket and are irreversible), bivalent inhibitors (binding the catalytic site and a second stability or localization site), allosteric inhibitors that bind distal regulatory sites (type III and IV), and degraders or molecular glues that hijack the ubiquitin–proteasome system to remove the kinase rather than inhibit it.
Concept Translation: Kinases are useful to keep in mind because they’re the canonical “well-modeled” target family - thousands of crystal structures, deep mutagenesis data, and a consistent pocket geometry across the family. They’re roughly the ImageNet of structure-based drug design: the family where everything was tuned first and where most ML benchmarks were established. The frontier targets are unlike kinases - flat surfaces, no obvious pocket, sparse training data - and that’s why the modality expansion matters.
Kinase degraders blur the classical taxonomy. A “kinase inhibitor” and a “kinase degrader” act on the same target through different mechanisms, so the druggability criteria differ as well. That overlap sets up the rest of this post.
Structure prediction and the 2020/2024 inflections
For most of drug discovery’s history, structure-based drug design was available only for targets that had been successfully crystallized. The c-Met/crizotinib program is a useful illustration of that earlier workflow. c-Met is a receptor tyrosine kinase with elevated levels and abnormal activity in several cancers. Pfizer’s program started from a series of pyrrole-substituted 2-indolinones (oxindoles), ATP-competitive tyrosine kinase inhibitors first explored in the mid-1990s, that showed activity against c-Met but poor drug-like properties. Iterative co-crystallography with PHA-665752 (an early lead) revealed the c-Met ATP-pocket environment in detail; that structure motivated a switch to a novel 2-amino-5-aryl-3-benzyloxypyridine series, which yielded the 3,5-disubstituted 2-aminopyridine clinical candidate crizotinib (PF-02341066, Cui et al., J. Med. Chem. 54:6342–6363, 2011). Each iteration of the design cycle was gated on a new crystal. Crizotinib, a c-Met/ALK dual inhibitor, was approved by the FDA in 2011.
That workflow still works, and for targets that crystallize well it remains the standard. Crystallography, however, is slow and expensive. For many targets - especially membrane proteins, intrinsically disordered regions, and transient complexes - it either fails or produces low-resolution artefacts that are hard to use for design.
Two inflection points changed the practical calculus.
2020: AlphaFold 2. At CASP14, the community benchmark competition for protein-structure prediction, DeepMind’s AlphaFold 2 produced predictions indistinguishable from experimental structures for a majority of single-chain proteins. By 2022 the AlphaFold Database had released ~800,000 predicted structures. By 2025 it was around 200 million predicted structures, covering the bulk of UniProt, the standard protein-sequence database. Single-chain structure prediction stopped being a bottleneck.
2024: AlphaFold 3. AF3 (Abramson et al., Nature 630:493–500, 2024) moved from single-chain prediction to complex prediction using a diffusion-based architecture built around a Pairformer block - which updates pairwise residue relationships - plus a diffusion module. It jointly predicts proteins with small-molecule ligands, DNA, RNA, ions, and post-translational modifications in a single forward pass. On the PoseBusters ligand-pose benchmark, a test of whether models place ligands in the right binding pose, DeepMind reports AF3 as roughly 50% more accurate than the best traditional physics-based docking tools. That pushes protein–ligand pose-prediction success rates to about 76% versus roughly 50% for the docking programs AutoDock Vina and Gold on the same benchmark, and makes AF3 the first AI system to outperform physics-based tools for biomolecular structure prediction. Model weights were released for non-commercial academic use in November 2024, with broader (still non-commercial) availability in February 2025. Commercial use remains gated behind Isomorphic Labs partnerships.
Concept Translation: The AF2-to-AF3 jump is conceptually familiar to anyone who has watched generative-model architectures evolve. AF2 was a transformer-style model that emitted a single deterministic structure. AF3 swaps the structure-emission head for a diffusion module - the same general framework as image diffusion models - that samples from a distribution over plausible structures. The Pairformer is a transformer-style block that operates on pairwise residue relations rather than tokens. The architectural pattern (encoder over pairwise relations + diffusion decoder) is now the template for every AF3-class model in the field.
The competitive field matters because AF3’s non-commercial licensing makes commercially usable alternatives important for industrial drug discovery:
- Boltz-1 (MIT, Nov 2024): MIT license, fully open source, matches AF3 accuracy on standard benchmarks. Boltz-1x added physics-based steering to reduce steric clashes; Boltz-2 (2025) added binding-affinity prediction, claiming roughly 2× the precision of standard ML and docking baselines on hit discovery.
- Chai-1 and Chai-2 (Chai Discovery): Apache 2.0 license, inference-only weights, can run with or without a multiple-sequence alignment. Chai-2 (2025) introduced zero-shot de novo antibody design with reported experimental hit rates around 16–20% on novel targets.
- Protenix (ByteDance, 2025), HelixFold-3 (Baidu), and OpenFold3 (in preview as of late 2025) round out the field.
By 2026, teams treat predicted structures as routine inputs to virtual screening and structure-based design. AF3-class models have substantially reduced the need for experimental crystallographic structures in early-discovery programs, particularly for targets where co-crystal structures are slow or expensive to generate. Virtual screening campaigns at 10K-compound scale now run in days on cloud GPUs.
A worked example: AlphaFold-enabled design for an undrugged target
A 2022 hepatocellular carcinoma (HCC) program provides the first published demonstration of using AlphaFold-predicted structures for hit identification in practice (Ren, Ding, Zheng et al., Chemical Science 14:1443–1452, 2023). The target, CDK20, also known as cell cycle-related kinase (CCRK), had no experimental structure and was effectively undrugged. Traditional structure-based design on CDK20 would have required first solving its crystal structure, a multi-year project in itself.
Instead, the team used the AlphaFold-predicted structure (AF-Q8IZL9-F1-model_v1) of CDK20 directly. The prediction was high-confidence across most of the protein except the C-terminus, which they removed because it occluded the solvent-exposed region of the ATP pocket. Pocket analysis identified a shallow ATP binding pocket of approximately 150 ų in the DFG-in conformation - one common active-state arrangement of kinase active-site residues - with MET84 as the hinge residue and PHE81 occupying the gatekeeper position (the hinge is where many kinase inhibitors make key hydrogen bonds; the gatekeeper residue helps control access to the back pocket). They fed the AlphaFold structure into Insilico Medicine’s generative chemistry platform (Chemistry42), which generated 8,918 candidate molecules. After docking, clustering, and pose inspection, seven were synthesized for the first round of testing. The first hit (ISM042-2-001) bound CDK20 with a Kd of 9.2 ± 0.5 µM (lower Kd means tighter binding) and was identified 30 days from target selection. A second optimization round produced ISM042-2-048, with Kd ≈ 567 nM and a CDK20 kinase-inhibition IC50 of 33.4 ± 22.6 nM - the concentration required to cut enzyme activity by half. ISM042-2-048 also showed selective antiproliferation against the CDK20-overexpressing HCC line Huh7 (cellular IC50 ≈ 209 nM) versus the HEK293 counter-screen (~1707 nM), a comparison cell line used to check selectivity.
AlphaFold converted a target that would previously have been deferred until crystallography succeeded into a target that could be designed against immediately. In programs that use this workflow, the structural-modeling portion of the pipeline has moved from years to weeks. Hit-to-lead optimization, preclinical pharmacology, and everything downstream still take the time they always did.
Compared with the crizotinib workflow described above, the difference is sharp. Crizotinib needed a crystal of each intermediate compound; CDK20’s lead compound was designed against an AI-predicted structure before any experimental structure existed. Structure-based drug design used to be gated on crystallography. In 2026 it is gated on whether the structure-prediction model is accurate enough at the specific pocket you care about. That is a softer gate, and the tooling keeps improving.
Beyond small molecules: modality choice is now a first-class question
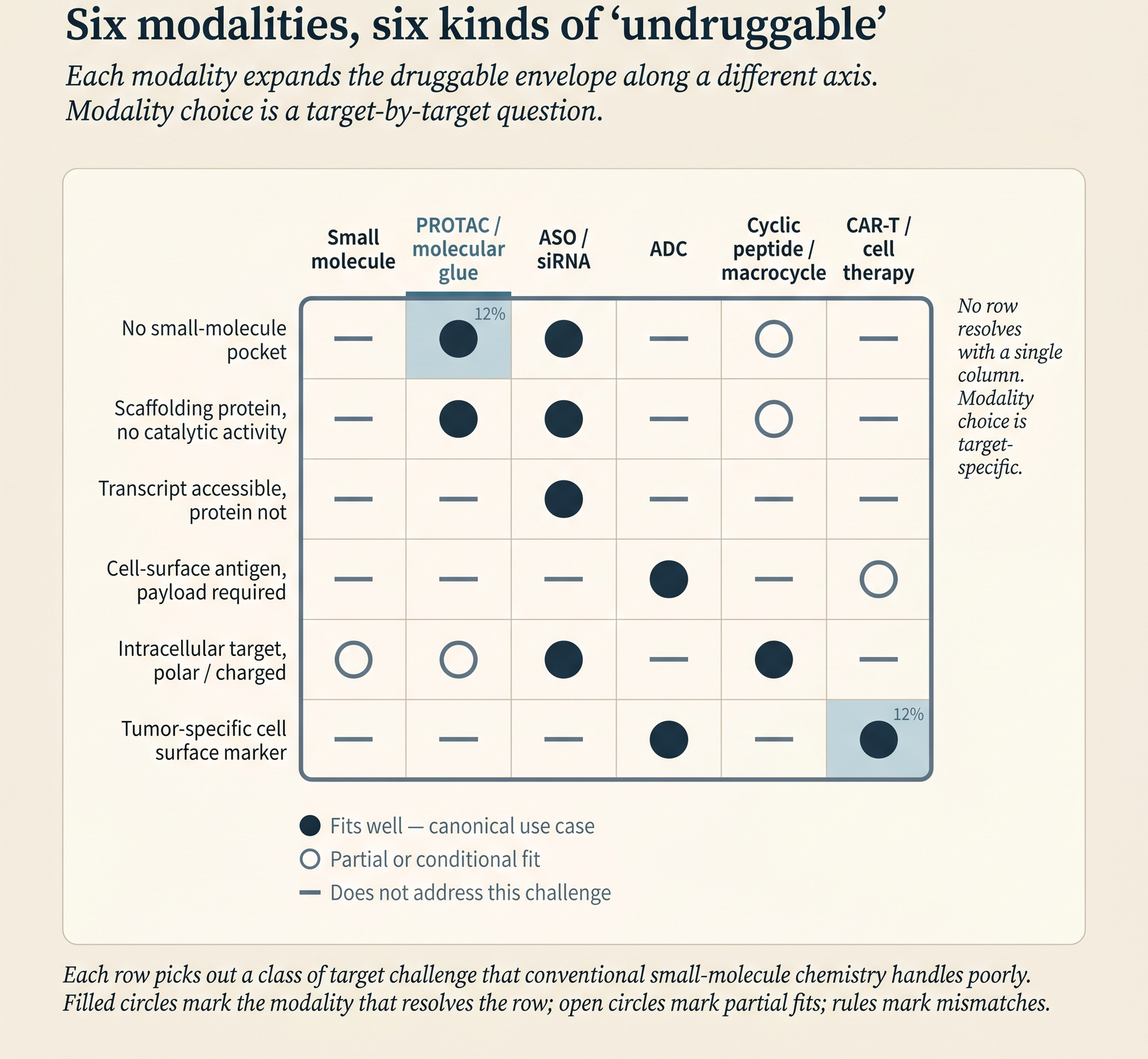
Categories of therapeutic molecule that did not have approved products a decade ago now have market-stage drugs and late-stage candidates, each reaching targets that small molecules cannot and expanding the druggable envelope.
Targeted protein degradation (PROTACs and molecular glues)
Targeted protein degradation (TPD) shifts pharmacology from occupancy-driven - where you block a protein’s function by occupying its binding site - to event-driven, where you induce the cell’s endogenous waste-disposal machinery to destroy the protein outright. The relevant machinery is the ubiquitin–proteasome system (UPS), the main pathway cells use to tag and remove unwanted intracellular proteins. A cascade of three enzymes - E1 (activating), E2 (conjugating), and E3 (ligase) - tags target proteins with polyubiquitin chains and marks them for destruction by the 26S proteasome. TPD drugs hijack this cascade.
Concept Translation: The occupancy-vs-event shift is closer to a control-flow change than a chemistry change. An occupancy-driven inhibitor is a function call that holds an object in a blocked state for as long as the call is on the stack - release the call, the object returns to normal. An event-driven degrader is a single instruction that triggers garbage collection on the object. The first scales with drug concentration; the second scales with the rate at which the cell remakes the protein. That difference is why PROTACs can be effective at much lower concentrations than equivalent inhibitors and why their pharmacology is often dominated by protein resynthesis rates rather than drug residence time.
A PROTAC (PROteolysis TArgeting Chimera) is a heterobifunctional molecule - it has one binding end for the target and another for an E3 ligase, joined by a linker. By forming a ternary complex (a three-part assembly of target, PROTAC, and ligase), it brings the target into proximity with the ubiquitination machinery. Because of this tripartite structure, PROTACs are large molecules, typically 700–1,000 Da. A PROTAC also acts catalytically. Once the target is ubiquitinated and released into the proteasome, the PROTAC dissociates and recruits another target copy, so one degrader molecule can remove many target molecules. It can therefore hit targets that have no functional small-molecule binding site at all, as long as some binder exists for the target.
Molecular glues are the close cousin: small molecules typically <500 Da, with no linker, that bind directly to an E3 ligase and induce a conformational change in the ligase surface, creating a novel interaction interface that recruits “neosubstrate” proteins (proteins the ligase would not normally bind) for degradation. The original IMiDs (thalidomide, lenalidomide, pomalidomide) are retrospectively understood as molecular glues. In 2026 the line between PROTACs and molecular glues is blurring; newer-generation degraders combine features of both.
The opportunity space is large. The human genome encodes over 600 E3 ligases, of which fewer than 2% have been successfully exploited for TPD to date. Most current PROTACs recruit just a few E3 ligases, especially CRBN or VHL; DCAF15 and MDM2 are emerging; the rest of the ligase repertoire is largely untouched. Tumor-specific E3 ligases are an active frontier for tissue-selective degradation. The motivation is direct: a CRBN-recruiting PROTAC can degrade its target in healthy tissue as well as diseased tissue.
The state of TPD in 2026 is a useful test case for “modality choice as a first-class question”:
- Bavdegalutamide (ARV-110), Arvinas’s androgen-receptor PROTAC, was the first PROTAC to enter human clinical trials (Phase 1, 2019). In patients with AR T878X/H875Y mutations in ARDENT, it produced 46% PSA50 response rates, where PSA50 is a prostate-cancer endpoint based on prostate-specific antigen decline. Development was discontinued in 2024 in favor of ARV-766, a next-generation AR degrader with broader mutant coverage. The first clinical entrant did not become the lasting lead program.
- Vepdegestrant (ARV-471), Arvinas/Pfizer’s estrogen-receptor PROTAC, reached Phase 3 (VERITAC-2) in estrogen-receptor-positive, HER2-negative breast cancer after prior CDK4/6 inhibitor failure. Topline results in March 2025 met the primary endpoint in the ESR1-mutant subpopulation, the subgroup whose tumors carried ESR1 mutations (median PFS 5.0 vs 2.1 months, HR 0.57, p<0.001). PFS is progression-free survival, and HR is the hazard ratio comparing event rates over time. The study did not reach statistical significance in the intent-to-treat population, meaning all enrolled patients. NDA, the New Drug Application, filed June 2025. PDUFA action date: June 5, 2026. If approved, vepdegestrant will be the first FDA-approved PROTAC.
- In September 2025, Arvinas and Pfizer publicly sought a third-party partner to commercialize vepdegestrant. That move suggests cooling internal commitment despite the positive result in the ESR1-mutant subgroup. The modality has clinical signal, but the commercial case remains unsettled.
The broader TPD space has expanded well beyond ER and AR. CC-94676 (BMS, AR PROTAC), the CELMoDs mezigdomide and iberdomide - cereblon E3-ligase modulators now in Phase 3 in multiple myeloma - and a growing number of Phase 1 molecular-glue degraders for BCL6, BTK, and other targets are all active programs.
For a target-assessment document, this changes the screen. For a scaffolding protein or transcription factor without an inhibitable catalytic site, the relevant question becomes “does it have a surface patch we can bind at all?” If yes, a PROTAC may be tractable.
RNA-targeting drugs (ASOs and siRNAs)
Antisense oligonucleotides (ASOs) are 15–30 nucleotide sequences that bind target RNA via Watson–Crick base pairing - that is, ordinary sequence complementarity. Two mechanisms dominate:
- RNase H1 cleavage. The ASO–RNA duplex is a substrate for endogenous RNase H1, an enzyme that cleaves the RNA strand. Most approved ASOs work this way.
- Steric blockage / splice switching. The ASO binds pre-mRNA (the RNA transcript before splicing) and prevents ribosomal assembly or alters splicing. Exon-skipping ASOs (Duchenne muscular dystrophy) and exon-inclusion ASOs (spinal muscular atrophy) are the canonical examples.
siRNAs act through the RNA-induced silencing complex (RISC), a cellular complex that uses the guide RNA to find and silence matching transcripts. Approved examples include patisiran (hereditary transthyretin amyloidosis), givosiran (acute hepatic porphyria), lumasiran (primary hyperoxaluria type 1), and inclisiran/Leqvio - Novartis’s siRNA targeting PCSK9, originally approved by the FDA in December 2021 for adults with ASCVD (atherosclerotic cardiovascular disease) or heterozygous familial hypercholesterolemia, expanded in 2023 to broader primary hyperlipidemia, and updated again in July 2025 to permit first-line monotherapy use in hypercholesterolemia.
Concept Translation: ASOs and siRNAs target messenger RNA rather than the protein itself. The matching is sequence-based - Watson–Crick base pairing is exact-match string lookup over a 4-letter alphabet (A, U, G, C). For target assessment this is a different design space entirely: the target is a string, the drug is a string, and the binding rule is a known function of the two strings. Compare this to small-molecule binding, where the binding rule is a learned function of two 3D shapes. Sequence-matching is much easier to design but introduces a new failure mode - off-target hits anywhere in the transcriptome that match closely enough.
For target assessment, ASOs and siRNAs require an accessible sequence rather than a structured protein binding pocket. A target with no deep pocket, no covalent warhead opportunity, and no PROTAC handle can still be tractable if its RNA is accessible. That rule has caveats: small-molecule splice modulators like risdiplam for SMA exploit RNA secondary structure - the local folded shape of the RNA strand - so structure does come back into play at the RNA level for some programs.
Antibody-drug conjugates (ADCs)
An ADC is a monoclonal antibody covalently linked to a cytotoxic payload via a chemical linker. The antibody provides selectivity; the payload provides potency. An ideal ADC stays stable in circulation, internalizes on binding to the target (the antibody–target complex is pulled into the cell), and releases the payload intracellularly in the vicinity of the target.
Concept Translation: An ADC is a delivery system. The antibody is a routing label that says “deliver to cells expressing this surface marker”; the payload is the actual cytotoxic drug, often too toxic to administer freely. The linker controls when the payload separates - ideally only after the package has been internalized. The pattern is roughly the same as targeted delivery in any system: route on a label, release on arrival. The engineering challenges are also analogous (label specificity, in-transit stability, controlled release).
2026 status: 15 ADCs are FDA-approved, with 19 ADCs approved globally across FDA, EMA, NMPA, and PMDA. Pipeline scale is over 400 ADCs in development, 200+ in clinical trials, and at least 24 in Phase 3. Recent FDA approvals include Datroway (datopotamab deruxtecan, January 2025) and Emrelis (telisotuzumab vedotin-tllv, May 2025, targeting c-Met-overexpressing NSCLC, or non-small cell lung cancer). Topoisomerase I inhibitor payloads (DXd, SN-38) now dominate late-stage development due to strong bystander effects, where the released drug can also kill neighboring cells.
For target assessment, ADCs open up targets where selective expression matters more than pocket tractability. A tumor-surface antigen that cannot be usefully modulated by a naked antibody (an antibody without an attached payload) can still be a good ADC target if it internalizes on binding.
Therapeutic peptides and cyclic peptides
Therapeutic peptides occupy the middle ground between small molecules and biologics. They typically fall in the 500–5,000 Dalton range, often offer high target specificity, are less likely to trigger an immune response than antibodies, and have better membrane permeability than most proteins. Linear peptides suffer from poor stability and short half-life. Cyclic peptides address both by constraining the peptide into a ring, which improves rigidity, stability, and permeability. Cyclic peptides sit within the broader “beyond-Rule-of-Five” (bRo5) chemical space - shorthand for compounds that break the usual small-molecule size and polarity heuristics. Macrocycles (large rigid rings of 12 or more atoms), PROTACs, peptides, and metallodrugs all populate this space. By 2024, the field broadly accepted that strict adherence to Lipinski’s Rule of Five was not a prerequisite for intracellular engagement, and that bRo5 chemistry was central to engaging the flat protein–protein interaction surfaces that dominate the “undruggable” set. Cyclic peptides have shown activity in oncology, antiviral, antibacterial, and antimalarial contexts.
CAR-T, gene therapy, cell therapy
CAR-T and TCR-T therapies genetically modify a patient’s T cells to express a chimeric antigen receptor or T-cell receptor, then infuse the modified cells back. Kymriah and Yescarta were the first two FDA approvals in 2017; Tecartus, Breyanzi, and Abecma followed for mantle cell lymphoma, large B-cell lymphoma, and multiple myeloma respectively. By early 2026, the FDA reported having approved close to 50 cell and gene therapy products over the prior decade across the Center for Biologics Evaluation and Research (CBER) Office of Therapeutic Products, the cumulative result of approvals such as Casgevy and Lyfgenia (sickle cell disease), Beqvez (hemophilia B), Elevidys (Duchenne muscular dystrophy), Vyjuvek (dystrophic epidermolysis bullosa), Aucatzyl (B-ALL, or B-cell acute lymphoblastic leukemia), and an ongoing run of CAR-T approvals.
For target assessment, CAR-T targets are a different class of object from small-molecule targets. A CAR-T target is a surface antigen expressed, ideally exclusively, on disease cells - the “druggability” question is about on-tumor versus off-tumor expression (whether the antigen appears on cancer cells, healthy cells, or both), not pocket geometry. That leads to the other half of the 2026 druggability question.
The druggable genome is expanding, and the ignorome remains large
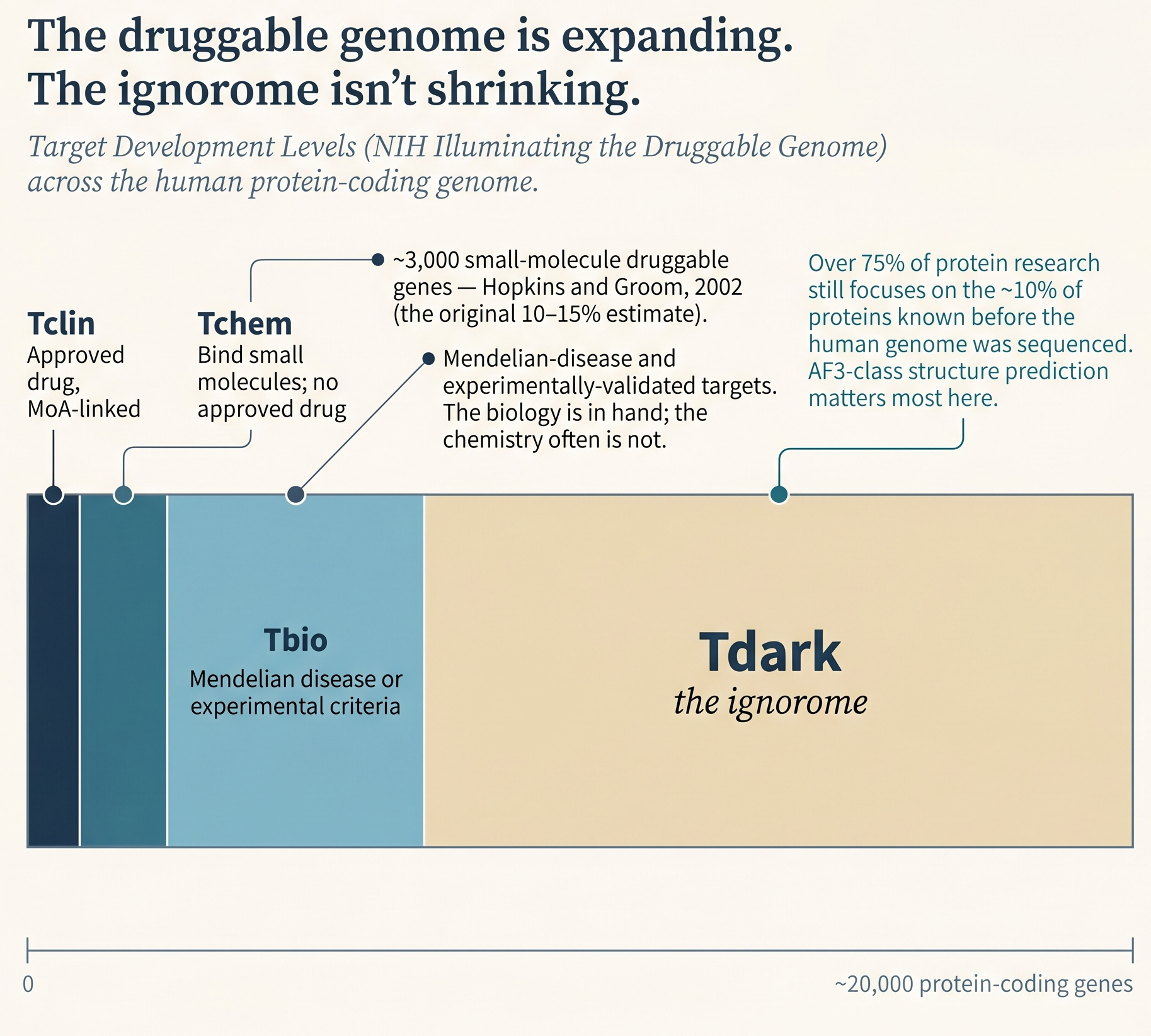
There are more than 10,000 known human diseases. The original 2002 estimate put 3,000–10,000 disease-related genes in the genome, with roughly 10% of those being disease-modifying on knockout (disabling the gene changed a disease-relevant phenotype in model systems). That yields the 600–1,500 small-molecule-druggable target estimate cited at the top of this post.
The Rule of Five (Lipinski et al., 1997) shaped small-molecule design for a generation: no more than 5 hydrogen-bond donors, 10 acceptors, molecular weight under 500, and LogP under 5, with LogP being a measure of lipophilicity. By 2026, approved drugs increasingly exceed Rule-of-Five boundaries. The main alternative is the Rule of Three for fragment-based discovery, a strategy that starts from very small weak-binding molecules (Congreve et al., Drug Discov. Today 8:876–877, 2003: MW < 300, ≤3 H-bond donors, ≤3 H-bond acceptors, cLogP ≤ 3, where cLogP is the computed version of LogP). The empirical boundaries of “drug-like” are shifting as chemistry expands to PROTACs, macrocycles, ADCs, and covalent warheads. These categories routinely break Lipinski’s original rules and still get approved.
Concept Translation: Rule of Five was a hand-crafted feature filter - a set of property thresholds that empirically separated absorbable drugs from non-absorbable ones, fitted on roughly 2,200 oral drugs in the late 1990s. Like any hand-crafted filter, it was tuned to its training distribution. As the chemistry expanded into larger and more polar molecules, the filter started rejecting things that worked. The 2026 stance is to use Lipinski thresholds as a soft prior for the small-molecule region of chemical space and to ignore them entirely outside it.
Meanwhile, over 75% of protein research still focuses on the 10% of proteins known before the human genome was mapped. The NIH’s Illuminating the Druggable Genome (IDG) initiative, launched in 2014, created a four-level Target Development Level classification to make this visible:
- Tclin: targets linked to at least one approved drug by mechanism of action.
- Tchem: proteins known to bind small molecules with high potency, but without approved-drug links.
- Tbio: proteins with a confirmed Mendelian disease phenotype or meeting certain experimental criteria.
- Tdark: proteins meeting none of the above.
IDG found that Tdark proteins receive less research funding than other categories, which perpetuates the knowledge gap. The ignorome - the Tdark set - is both the largest opportunity space for novel target discovery and the hardest place to work, because almost every downstream tool (antibodies, assays, knockout models) has to be built from scratch. The computational tractability gains from AF3-class structure prediction matter most here: a target with no crystal structure and minimal biological characterization now at least comes with a predicted structure to work from.
Concept Translation: The TDL classification is a label hierarchy on the human proteome - closer to a knowledge-graph annotation than a clean training-data split. Tclin proteins have the most data on them (drugs, structures, papers, assays) and Tdark have the least. ML methods trained on protein-level features inherit the same skew as the labels: they perform best on Tclin and worst on Tdark, the long tail. The “ignorome” framing emphasizes that the tail is where novel drug targets actually live.
A 2026 assessment of a novel target should include, at minimum, an AF3-class structure prediction with pocket analysis, a TDL classification, a scan across modality options (small molecule, PROTAC, ASO, ADC, peptide), and an explicit note on whether Rule-of-Five chemistry is even the right design envelope for this target. The question [novelty scoring and the ignorome] → Novel vs repurposed targets: quantifying novelty and extending drug-repurposing methods (C5)1 takes up how novelty itself is now quantified across the TDL spectrum.
What’s next in druggability assessment
By 2026, the central druggability question is “which of five or six modalities fits this target best, and what is the safety profile of hitting it that way?” Three follow-ups stand out.
Safety. A PROTAC that catalytically degrades its target in tumor tissue may also degrade the same target in healthy tissue where it is essential. An ASO that knocks down a transcript in the liver may behave differently in the kidney. Modality-specific on-target, off-tissue toxicity - where the intended target is hit in the wrong tissue - is the next frontier, and it’s directly continuous with the tissue-specificity question the next post in this series picks up: [tissue specificity] → Tissue specificity as a safety filter: GTEx, Human Protein Atlas, and scRNA-seq for target prioritization (C3).2
Two targets, not one. The entire framing of druggability assumes you’re choosing a single target and asking what modality hits it. For some diseases, most notably cancers with defined synthetic-lethal vulnerabilities (where a cell tolerates either perturbation alone but not both together), the right answer is that you need to hit two targets together, and each of those targets individually is a poor drug candidate. That shifts the druggability question to a two-body problem, with its own computational and experimental methods: [synthetic lethality] → Synthetic lethality and combination targets: ML methods for finding drug pairs that work together (C7).3
Benchmarks for modality-choice models. We don’t yet have a standard benchmark for “given this target, which modality will work best?” The PoseBusters benchmark answers a narrower question (given this target and this ligand, is the predicted pose right?). Benchmarks for PROTAC ternary-complex geometry, ASO off-target profiles, and ADC internalization kinetics exist in pieces but do not yet form a coherent evaluation suite. That gap is a natural target for the next round of infrastructure investment. For the pillar-post view of where this fits in the overall workflow: [the pillar post on drug target discovery] → Target discovery: the front-of-funnel decision behind most Phase II failures (P0).
AlphaFold 3 was an inflection point. The remaining work now sits in modality selection, safety, and benchmark design.
Further reading
This Further Reading section leans on primary papers rather than review-heavy framing because the topic dates quickly. AF3 is two years old at the post’s publication, PROTAC/TPD clinical status moves every few months, and the ADC field adds approvals on a rolling basis. Primary papers and canonical databases age more gracefully.
-
Abramson, Adler, Dunger, Evans, et al. (2024). Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 630, 493–500. DOI: 10.1038/s41586-024-07487-w. The primary source for every AF3-specific claim in the post, including the PoseBusters benchmark, the Pairformer-plus-diffusion architecture, and the joint protein–ligand–DNA–RNA prediction capability. Cited rather than a more recent review because the benchmark numbers in reviews inherit from this paper anyway. The competing AF3-class models (Boltz-2, Chai-2, Protenix, OpenFold3) have released benchmark head-to-heads since 2024; readers building a current-day pipeline should consult the relevant tool’s release notes for the most up-to-date head-to-head numbers.
-
Hopkins, A. L., and Groom, C. R. (2002). The druggable genome. Nature Reviews Drug Discovery 1, 727–730. The foundational paper that anchors the “10–15% of the proteome is druggable” framing used throughout the post’s definitional section. Oprea, Bologa, Brunak et al. (2018), Unexplored therapeutic opportunities in the human genome, Nature Reviews Drug Discovery 17, 317–332, is the closest-to-current methodological update and provides the Tclin/Tchem/Tbio/Tdark classification that the ignorome section discusses - consider citing it alongside Hopkins & Groom if the post needs a more current primary reference.
-
Ren, F., Ding, X., Zheng, M., et al. (2023). AlphaFold accelerates artificial intelligence powered drug discovery: efficient discovery of a novel CDK20 small molecule inhibitor. Chemical Science 14, 1443–1452. DOI: 10.1039/d2sc05709c. The primary source for the worked CDK20/HCC example used in the post - the first published demonstration of AlphaFold-predicted structures driving hit identification end-to-end. Verifies the 30-day target-to-hit timeline, the 8,918-molecule generative pool, the MET84/PHE81 hinge–gatekeeper geometry, and the ISM042-2-048 33 nM IC50 figure.
-
Cui, J. J., Tran-Dubé, M., Shen, H., et al. (2011). Structure based drug design of crizotinib (PF-02341066), a potent and selective dual inhibitor of mesenchymal–epithelial transition factor (c-MET) kinase and anaplastic lymphoma kinase (ALK). Journal of Medicinal Chemistry 54, 6342–6363. DOI: 10.1021/jm2007613. The medicinal-chemistry case history behind the crizotinib comparison - documents the path from pyrrole-substituted oxindoles through PHA-665752 to the 2-aminopyridine clinical candidate, including the iterative crystallography that the AF3-era CDK20 workflow makes optional.
-
NIH Illuminating the Druggable Genome Program. Pharos: the IDG Knowledge Management Center’s public resource for target development-level classification. pharos.nih.gov. The canonical open-access source for the TDL classification (Tclin/Tchem/Tbio/Tdark) the post uses in its ignorome section. Cited as the database home page rather than a specific IDG paper because the TDL categories are continuously updated against new drug approvals and omics evidence - a release-notes or current-database link ages better than a one-time publication link.