Drug target discovery: the front-of-funnel decision behind most Phase II failures
A drug candidate can enter Phase II clinical trials with a nine-figure budget behind it, years of preclinical work completed, and an IND (Investigational New Drug application) cleared by the FDA, yet still carry the wrong answer to a question asked five to ten years earlier: is this the right biological target? For readers new to the pipeline, Phase I is mainly about safety and dosing. Phase II is where efficacy first gets a serious test in patients. Phase III is the larger confirmatory stage. Roughly 72% of drugs entering Phase II do not transition to Phase III, and approximately 90% of investigational drugs fail somewhere in clinical development overall (source: GlobalData clinical analytics, 2024). Lack of efficacy against the intended disease accounts for roughly half of all clinical trial failures; another ~30% are halted for unmanageable safety and toxicity findings. In many cases, the problem starts before dosing or formulation matter. The target was wrong, or the evidence for its role in the disease was too weak. Target discovery is one of the most consequential stages of the drug development pipeline, and this series intends to teach why.
Why drug target discovery matters in 2026
When it comes to understanding chemistry and biology, the models are getting better. Generative models can design and optimize novel small molecules with greater success. AlphaFold 3 and the open-weights field around it, meaning models released with their parameters, have made routine structure prediction a solved problem for many classes of proteins. Docking, virtual screening, and ADMET prediction (absorption, distribution, metabolism, excretion, and toxicity) continue to mature as well.
Yet clinical success rates have worsened. Industry-wide likelihood of approval (LOA) for a Phase I compound fell from about 10.4% for the 2014 single-year cohort to around 6.7% (average LOA for 10-year period from 2014-2023). Phase II remains the dominant attrition step, with only about 28% of drugs that enter Phase II making it to Phase III (phase-by-phase transitions: Phase I = 47%, Phase II = 28%, Phase III = 55%, Filing-to-approval = 92%).
To assist instruction, front-of-funnel ML now has a concrete clinical example. In June 2025, Nature Medicine published Phase IIa results for rentosertib, a TNIK inhibitor for idiopathic pulmonary fibrosis (IPF), whose target (identified from multi-omics analysis of IPF patient tissue, meaning joint analysis of several molecular data types) and molecule (generative chemistry) were both AI-derived end-to-end. The study was small and short: 71 patients over 12 weeks (it was hypothesis-generating rather than a study designed to support approval) with reported improvements in standard lung-function measures relative to a placebo. Still, it marked the first time a fully AI-discovered drug, target included, produced a direction-of-effect efficacy signal in humans. Industry trackers now count more than 173 AI-discovered drug programs in clinical testing as of early 2026.
This post lays out the series: what target discovery is, why it matters, what makes it hard, and where ML helps. Each subsequent post will take one axis of target discovery and examine it in detail:
- How to tell a drug target matters using evidence frameworks
- Druggability, ligandability, and modality choice in the AlphaFold 3 era
- Tissue specificity: the safety half of target selection
- The most crowded and abandoned therapeutic areas
- Novelty vs repurposing: when to invent a target and when to reuse one
- Synthetic lethality and combination targets: ML methods for finding drug pairs that work together
- Knowledge graphs and case studies in AI-driven target discovery
- Virtual cells for target discovery, perturbation models, and benchmarks
What a drug target is
A drug target is a biomolecule, usually a protein and sometimes a nucleic acid, whose activity we want to modulate to produce a therapeutic effect. “Modulate” covers a range: inhibit an overactive enzyme, block a receptor from binding its ligand, degrade a disease-driving protein, replace a missing transcript. Binding the target has to produce the therapeutic effect. A molecule that binds to the target without changing disease outcomes is hitting a decoy.
Nearly half of oral drugs on the market target enzymes, with kinases as the single most productive sub-family. About a third target cell-surface receptors, especially G-protein-coupled receptors. Ion channels, transporters, and nuclear hormone receptors round out the major protein classes. Nucleic acids form a smaller but fast-growing non-protein class. If you’ve read chapter 1 of Machine Learning for Drug Discovery, we derive a ~10⁵ protein “biological search space” covering on the order of 100,000 potential human protein targets once splice variants and post-translational modifications, that is, alternative versions of a protein and chemical changes added after it is made, are counted. A drug-like molecule has to find, bind, and modulate the right one.
Target discovery, the front-of-funnel activity this series focuses on, is the work of deciding which of those proteins to go after for a given disease. It splits into two phases that casual usage often folds together:
- Target identification: generating candidate target-disease links. “Here are ten proteins that seem to be involved in pancreatic fibrosis; let’s prioritize the top three.”
- Target validation: building the evidence that one of those candidates is actually causal. Does knocking it out in a disease model reverse the phenotype? Does a human genetic signal support it? Is there pharmacological precedent?
Both phases happen before a single compound is screened. Get either wrong and the downstream pipeline is solving the wrong problem.
Funnel Economics: The Cost of Being Wrong at Target Selection
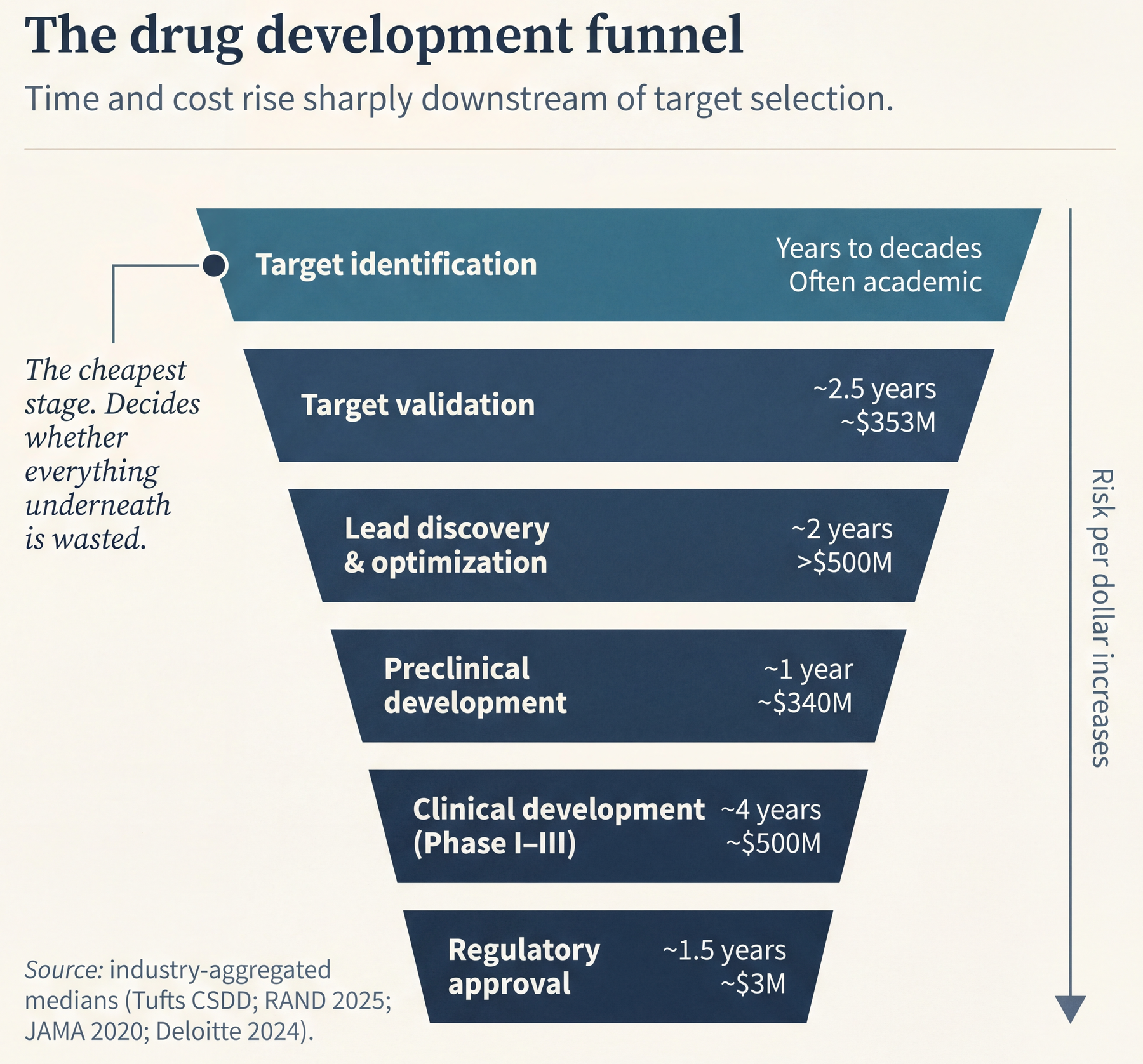
The drug-development funnel runs roughly like this:
- Disease hypothesis and early target identification: years to decades, often academic.
- Target validation: up to ~2.5 years, averaging around $353M when done end-to-end including follow-on studies.
- Lead discovery and optimization: up to ~2 years, over half a billion dollars.
- Preclinical development: about a year, ~$340M.
- Clinical development: Phase I about 1.5 years, Phases II/III about 2.5 years combined, roughly half a billion dollars in total.
- Regulatory approval: ~1.5 years, ~$3M, the cheapest transition once you’re there.
Aggregate “cost to develop a new drug” figures vary widely because analysts treat time costs, failure allocation, and capitalization differently. The Tufts Center for the Study of Drug Development’s widely cited $2.8 billion mean per-approved-drug cost includes roughly $1.16 billion of foregone-investor returns over the development window plus $312 million in post-approval R&D. More recent cost estimates from RAND (2025), JAMA (Wouters et al., 2020), and Deloitte (2024) report different distributions, with medians closer to $0.7 billion to $1.0 billion and means weighted by high-cost outliers in the $0.95 billion to $2.23 billion range. Though reported headline figures depend heavily on which costs are included, the point is consistent. This is an expensive process, and being wrong costs a lot of cash!
That makes the target-selection decision unusually consequential. It spends comparatively less money, yet determines whether everything underneath it is wasted. Successful or unsuccessful completion of Phase 2 for an individual drug costs pharmaceutical companies around 20% of the sum spent on the drug discovery pipeline recalculated for each individual drug. Phase III is a larger absolute commitment, but Phase II is where we first get an answer to “is this target right?” and where many disappointments occur.
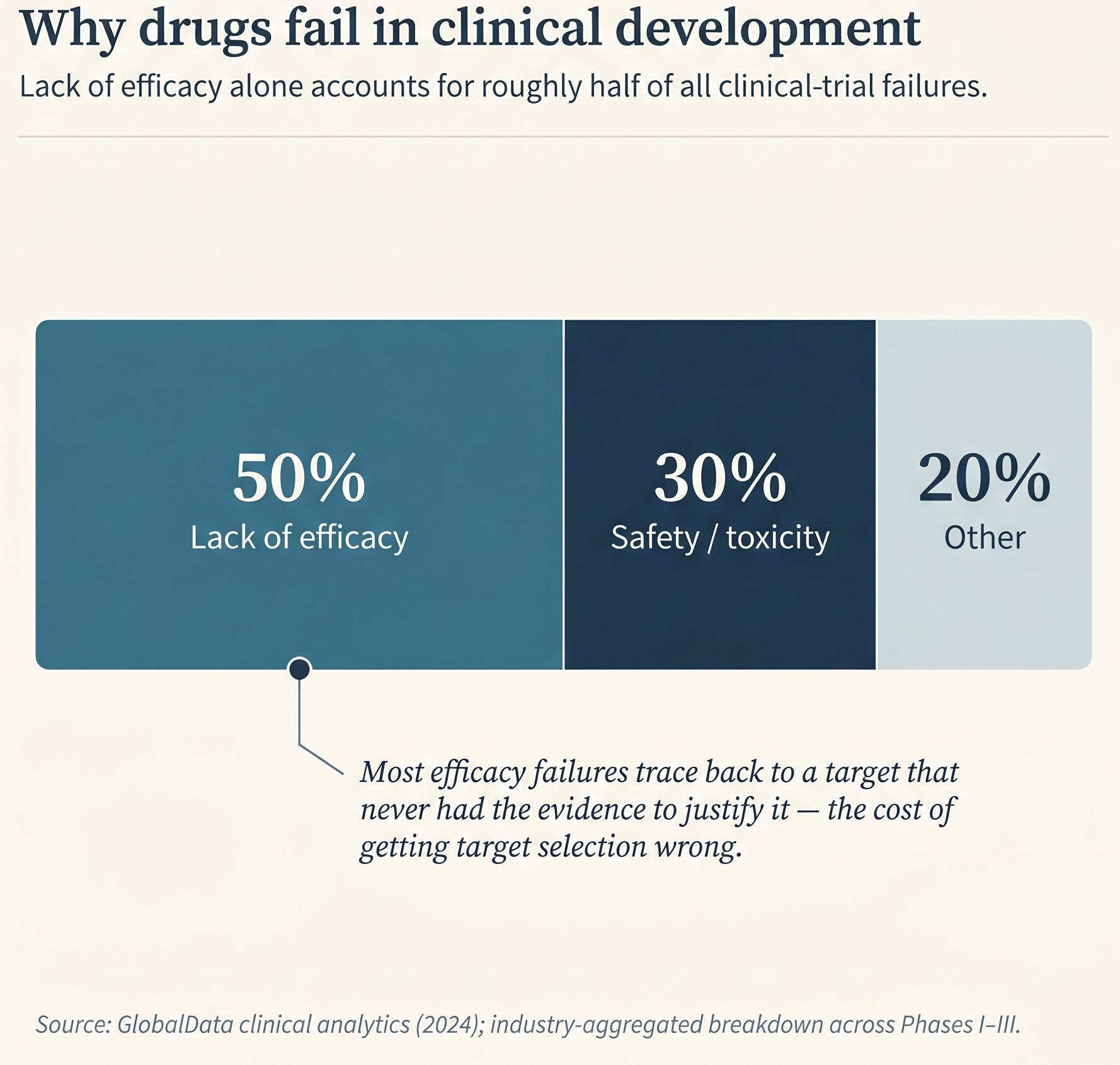
In a 2024 report, GlobalData attributes roughly half of clinical trial failures across phases to lack of efficacy and another ~30% to safety or toxicity. If efficacy collapses because the chosen target never had enough evidence behind it, then better target selection compounds downstream. Prevent one Phase II failure and the savings can exceed what you spent on target-assessment work by an order of magnitude. Catch the same mistake at target selection instead of in Phase II and the savings are larger still.
The 2014–2023 industry data also points to a second problem. Phase I likelihood of approval fell from 10.4% for the 2014 cohort to 6.7% for the most recent ten-year cohort, partly driven by an industry shift toward first-in-class, riskier targets where the evidence base is thinner. Many easier targets have already been drugged. What’s left is harder, and harder problems put more weight on front-of-funnel methods. Biomarker-stratified programs, which enroll patient subgroups defined by a measurable biomarker, run at roughly double the LOA of unstratified ones. This result follows from stronger target-selection evidence, as the biomarker identifies the patient subgroup in whom the target matters. For this reason, target and biomarker development often move together.
Why drug target discovery is hard
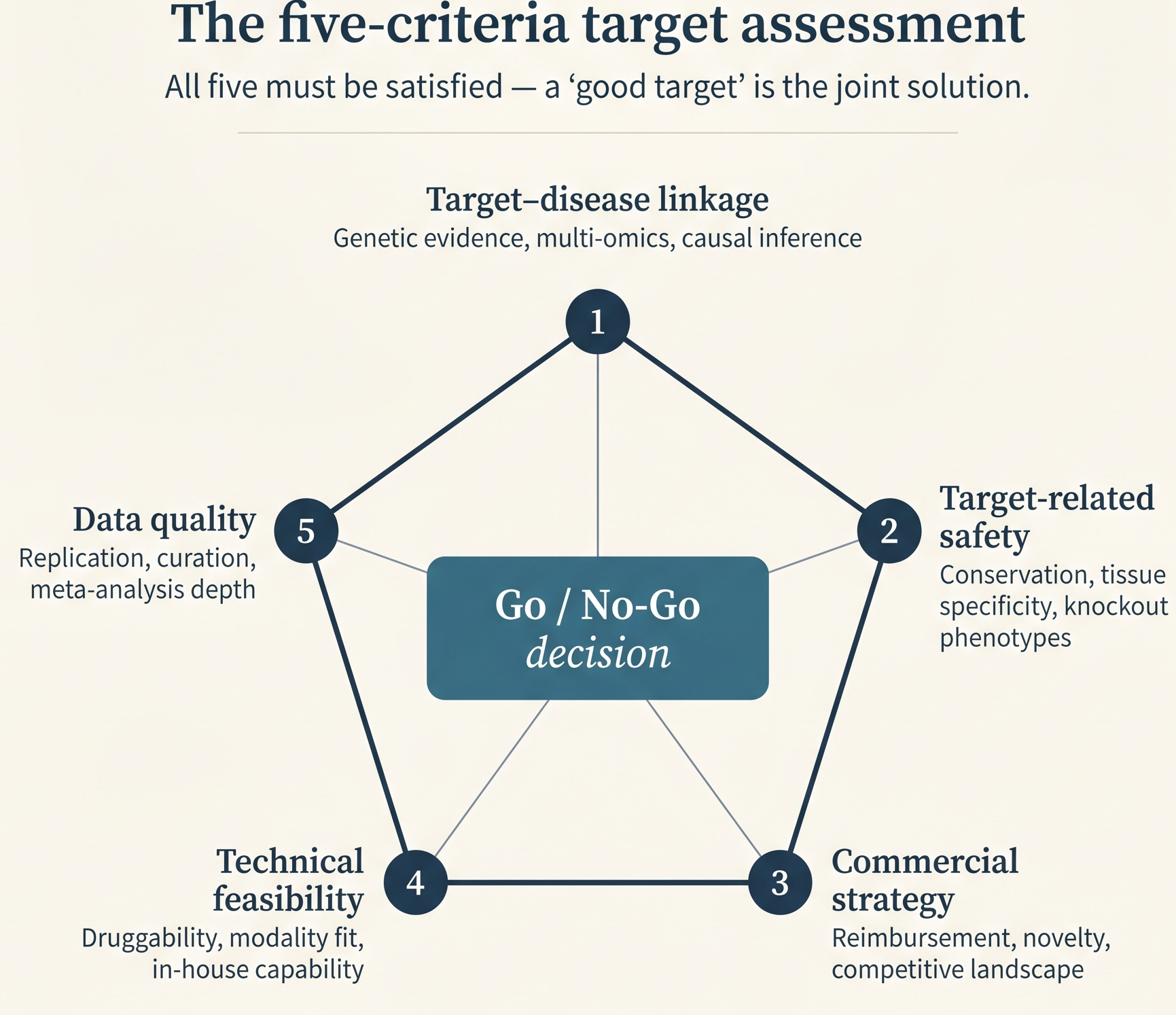
There is no single solution or approach as every target hunting strategy is context dependent. Useful target-assessment work has to satisfy at least five criteria simultaneously.
Target–disease linkage
Does the target play a causal role in the disease rather than merely correlate with it? Genetic evidence carries the most weight in most organizations. That includes GWAS hits, population-scale studies that link genomic loci to disease, Mendelian randomization, which uses inherited variants as a quasi-natural experiment, and rare-variant burden tests, which ask whether damaging variants in a gene accumulate more often in cases than controls. Multi-omics integration, which combines several molecular data layers such as RNA, protein, and chromatin measurements, acts as supporting evidence.
Target-related safety
Can humans tolerate modulation of this protein for years? Evolutionary conservation predicts essentiality and toxicity, wwhereas tissue specificity predicts off-target effects. Knockout phenotypes also provide a useful preview into safety implications if we were to neutralize the target.
Commercial strategy
Is there a path to reimbursement? Is this a first-in-class program, meaning a new mechanism, a best-in-class program, meaning a superior drug against an established mechanism, or a repurposing play, meaning an existing drug or target used in a new disease? These questions inform which target to prioritize even when several are scientifically equivalent.
Technical feasibility
Is the target druggable, meaning some therapeutic class can plausibly bind or modulate it? Can the organization develop the right modality, such as a small molecule, antibody, or oligonucleotide? A protein-protein interaction with no obvious pocket may be validated yet undruggable.
Data quality
Is the evidence base solid, or built on a few underpowered studies that never replicated? Manual curation, field-standard evaluation, and in some cases meta-analyses feed into this.
Taken together, a “good target” satisfies a multi-objective optimization problem across all five properties, involving a variety of data modalities. In practice, these criteria produce weighted votes rather than verdicts. Few targets get a clean yes across all five. More often, we see mixed signals and have to decide whether one target candidate’s profile beats the other eight candidate targets on our list. Target-discovery ML tries to improve this decision-making process.
Where ML helps in drug target discovery
The chemistry-side ML playbook of generating candidates, predicting properties, and optimizing molecules does not transfer cleanly to target discovery. There is not an explicity accuracy metric to optimize against, training data is fragmented across a dozen modalities, and ground truth, i.e., “was this the right target?”, takes a decade and often a Phase III trial to establish. Despite these limitations, ML does three things well here.
Scale
Datasets for target discovery are enormous and heterogeneous:
- GenBank contains over 300 billion base pairs of sequencing data.
- UniProt has more than 180 million protein sequences.
- The ENCODE multi-omics database exceeds one petabyte.
- PubMed has more than 35 million citations
- The USPTO database has hundreds of thousands of patents
- ClinicalTrials.gov has more than 438,000 clinical studies
No human team can read or integrate all of this. ML can convert it into searchable, queryable representations. Knowledge graphs built from biomedical text, which turn papers into linked entities and relationships, compressed numerical representations of multi-omics datasets, and network-proximity algorithms that rank genes by their topological distance from disease-associated nodes are all methods that scale to the size of the actual evidence base.
Cryptic patterns
A well-known example comes from SARS-CoV-2 repurposing, where we consider existing drugs that might work in a new or unaddressed disease. One study used network-diffusion and network-proximity algorithms on a combined dataset of the human interactome, the network of protein-protein interactions in cells, viral targets, and drug interaction data to rank 6,340 drugs for expected efficacy. The top-ranked drugs, tested experimentally, showed a 62% success rate in reducing viral infection, and 76 of the 77 drugs that worked did not bind to proteins directly targeted by the virus, suggesting network-based mechanisms rather than direct target engagement. That kind of signal, which requires multiple hops of complex reasoning, is hard to recover by manual review. In this case, graph algorithms operating across the interactome were able to surface it.
Automation
Literature review for a new target can take days or months. Named-entity recognition, which identifies mentions of genes, diseases, drugs, authors, and institutions in text, can be automated with usable quality (state-of-the-art NER reaches roughly 90% F-score). Relation extraction, which asks whether a paper asserts a specific link between those entities, is harder, around 50%. A knowledge graph that updates as new papers are published treats the evidence base as a living object, instead of freezing it at the last manual review.
These are front-of-funnel tasks that ML handles well. Final prioritization is a different matter*. In practice, target selection remains heavily human-curated. ML produces ranked candidate lists, aggregated evidence dashboards, and network-scored hypotheses, but the decision to allocate a program’s budget to target #3 rather than #7 still depends on organizational expertise, portfolio balance, competitive intelligence, and the intellectual-property landscape. Those factors are usually absent from training data. For now, target-discovery ML compresses the evidence into a form a human committee can reason about.
*If you’re coming from a background in recommender systems, this is similar to a cascading systems approach where we care about maximizing recall at the start (i.e., we don’t want to preemptively filter out a candidate and cut a potential multibillion dollar revenue stream) and, by the end, are most trying to maximize precision (i.e., avoid a costly false positive that damages our corporate brand, wallet, and sanity).
Worked Example: Rentosertib
Insilico Medicine’s rentosertib (also known as ISM001-055 / INS018_055) for idiopathic pulmonary fibrosis (IPF) serves as an instructional walkthrough (though I do not go so far as to categorize it as a gold-standard benchmark) of one end-to-end AI-driven target-and-molecule program.
Target identification began with multi-omics analysis of lung tissue from IPF patients versus healthy controls, combined with text mining across IPF literature and knowledge-graph reasoning over protein-protein interactions and pathway data. TNIK, a TRAF2- and NCK-interacting kinase, emerged as a novel regulator of fibrotic pathways. I.e., TNIK was not an established IPF target when the program started. The timeline is also notable. Project initiation to preclinical candidate nomination, the point where a program chooses the molecule it will advance into formal preclinical testing, took roughly 12–18 months, and the asset reached human clinical trials in under 30 months total, against an industry average of 4.5–6 years for early-stage discovery. Target validation ran in parallel with early chemistry; preclinical models confirmed that TNIK inhibition reduced fibrotic phenotypes, and generative-chemistry tools designed a series of inhibitors. The knowledge-graph workflow behind this will be the subject of a future article in this series.
Phase I established safety and pharmacokinetics, meaning how the body absorbs, distributes, and clears the drug. The Phase IIa trial (GENESIS-IPF, NCT05938920) enrolled 71 IPF patients across 21 sites in China over 12 weeks. The secondary endpoint, change in forced vital capacity (a standard measure of lung function) from baseline at week 12, showed +98.4 mL for the 60 mg daily dose versus −20.3 mL for placebo. Nature Medicine published the results in 2025, where they were positioned as the first clinical proof-of-concept for an end-to-end AI-discovered drug.
Keeping limitations in view, seventy-one patients over 12 weeks is a hypothesis-generating dataset, not a study sized or designed to support approval. No pivotal trial is running as of April 2026; Insilico is in regulatory discussions about a Phase IIb pivotal study, and a separate US Phase IIa (NCT05975983) is enrolling, with eight of the planned 60 patients having completed the 12-week treatment as of mid-2025. Keeping within the subject matter of this article series, TNIK had to be identified as a target before any chemistry could start. That identification came from a workflow combining multi-omics data, literature mining, and knowledge-graph reasoning. The Phase IIa result matters because it is consistent with the target call having been right. Whether the methodology generalizes is a question the next decade of readouts will answer.
A useful counterweight is Recursion’s REC-994, a lead pre-merger AI-discovered candidate for cerebral cavernous malformation, which was discontinued in May 2025 after long-term Phase II data failed to confirm earlier efficacy trends. High-profile AI-guided programs can fail in the clinic for the same reason other programs do. Through curating better evidence, we hope to lower the failure rate.
Who does drug target discovery
For readers whose mental model of drug discovery centers on Big Pharma, it’s motivating to keep in mind that, in the United States, nearly 60% of newly approved drugs were discovered in universities or biotechnology companies, not by Big Pharma’s own R&D. Small, often academic-adjacent biotechs take the early-stage innovation risk. Big Pharma licenses, acquires, or partners to bring the late-stage program through approval and marketing.
The reasons are structural. Big Pharma faces the “better than the Beatles” problem (the bar for new drug approval keeps rising because better drugs already exist), the “low-hanging fruit” problem (the easy targets are mostly drugged), the “cautious regulator” problem (FDA standards ratchet up after each safety scare and rarely relax), and a tendency to industrialize the wrong activities; scaling basic research and brute-force screening has not improved clinical success rates in aggregate. For more information, we discuss these problems and the related Eroom’s Law in detail within chapter 1 of “Machine Learning for Drug Discovery.”
For a practitioner in 2026, this means target-discovery tooling is disproportionately built inside smaller, often AI-native companies, licensed into Big Pharma programs, or run academically against public data. Big Pharma’s target-discovery groups increasingly act as evaluators and integrators rather than primary generators.
The rest of the drug target discovery series
The rest of this series goes deep on what this pillar surveys:
- Evidence frameworks → How to tell a drug target matters Driver vs passenger mutations, oncogene addiction (tumors becoming unusually dependent on one gene), multi-omics integration, GWAS, and practical target-assessment frameworks such as AstraZeneca’s 5R and GOT-IT.
- Druggability in the AlphaFold 3 era → Druggability, ligandability, and modality choice in the AlphaFold 3 era Classical druggability, the expanding druggable-genome concept, therapeutic modalities such as antisense oligonucleotides (ASOs), PROTAC degraders, and antibody-drug conjugates (ADCs), and what AlphaFold 3 changed about structure-based drug design.
- Tissue specificity → Tissue specificity: the safety half of target selection Single-cell RNA-seq and reference atlases such as GTEx and the Human Protein Atlas, and how tissue-level gene-expression data feeds into safety prediction.
- Likelihood of approval by therapeutic area → The most crowded and abandoned therapeutic areas Likelihood of approval by therapeutic area, the economics of repurposing, and why some disease areas are systematically more tractable.
- Novelty vs repurposing → Novelty vs repurposing: when to invent a target and when to reuse one When to invent a new target and when to reuse one. The Illuminating the Druggable Genome program, which focuses on understudied proteins, and research bias toward well-studied proteins.
- Synthetic lethality → Synthetic lethality and combination targets: ML methods for finding drug pairs that work together CRISPR-based screening, synthetic lethality, where dual perturbation of two genes kills a cell even though either single perturbation does not, and ML methods for drug-synergy prediction, including tools such as MAGeCK, CRISPRi, and Perturb-seq.
- Knowledge graphs and the rentosertib case study → Knowledge graphs and case studies in AI-driven target discovery Biomedical named-entity recognition, relation extraction, knowledge-graph embedding models, and walkthroughs of publicly documented AI-discovered-drug programs.
- Virtual cells → Virtual cells for target discovery, perturbation models, and benchmarks Single-cell atlases, perturbation-response models, Perturb-seq benchmarks, and how simulated interventions can prioritize target-validation experiments.
How drug target discovery connects to Machine Learning for Drug Discovery
Machine Learning for Drug Discovery concentrates on methods that begin after a target has been chosen. Molecular property prediction, virtual screening, generative chemistry, protein structure prediction, drug repurposing, and multimodal pipelines are chapter-length topics because they are well-defined ML problems with benchmarks and data.
If you came here from the book and have already built a property predictor, screened a virtual library, or trained a generative model for lead compounds, this series asks a prior question: how did anyone decide that was the right target in the first place? As you read through this series, you might notice that the methods change with the problem. For example, the work may involve graph learning over biomedical knowledge graphs, natural-language processing over biomedical literature, or exploiting multi-omics integration with multimodal models. However, at their core, these methods are more like variants of the methods we discuss in the book, rather than new or alien concepts.
What’s next in drug target discovery: open questions
Near-term readouts to watch. The 15 or so AI-discovered programs expected to enter pivotal Phase III trials in 2026 will provide the first broad test of whether front-of-funnel AI improves late-stage success rates. Rentosertib is the clearest test of AI-led target identification, because the target-selection step itself was AI-derived. Schrödinger/Takeda’s zasocitinib (TYK2 inhibitor) is a corresponding test of physics-based AI design.
Methodological open questions. First, how do you benchmark target-discovery ML without waiting a decade for each label? Surrogate endpoints, or proxy benchmarks, such as asking whether a method can rediscover known targets for known diseases using only pre-discovery literature, help but do not close the loop and may suffer from data leakage, data snooping, and related problems. Second, how far does the current generation of target-discovery ML extend beyond well-studied indications? The “ignorome,” the large set of human proteins that remain thinly studied, biases the training data toward what has already been studied. A target-discovery method that mostly finds targets adjacent to known ones misses the hardest part of the problem. The ignorome is the subject of a future article on novelty vs repurposing.
Regulatory shift. The FDA’s January 2025 draft guidance on AI in drug development established a risk-based credibility framework for when and how AI contributions must be qualified for regulatory submissions. Final guidance is expected within 2026, which will influence how target-discovery AI can be cited in an IND package (the dossier submitted to request permission for human testing).
For the first time, the ML methods used in target discovery are starting to produce clinical readouts. However, the jury’s still out and the next decade will show how much they change outcomes – anyone telling you differently has a financial interest competing with transparency ;)
Further reading
-
Citeline / Norstella (2024). Why are clinical development success rates falling? Industry report. Norstella/Citeline article. The primary source for the 6.7% Phase I LOA figure and the phase-by-phase transition rates cited throughout this post.
-
Sun, D. et al. (2025). A generative AI-discovered TNIK inhibitor for idiopathic pulmonary fibrosis: a randomized phase 2a trial. Nature Medicine. doi:10.1038/s41591-025-03743-2. The rentosertib Phase IIa paper that the “worked example” section walks through; the primary source for readers who want to see the trial design, endpoints, and safety data in full.
-
FDA Center for Drug Evaluation and Research (2025). Considerations for the Use of Artificial Intelligence to Support Regulatory Decision Making for Drug and Biological Products. Draft guidance, January 2025. FDA draft guidance. The regulatory framework being written for how AI contributions - including target-discovery AI - must be qualified in IND and NDA submissions, that is, submissions to begin human testing or request approval.
-
Wouters, O. J., McKee, M., Luyten, J. (2020). Estimated Research and Development Investment Needed to Bring a New Medicine to Market, 2009–2018. JAMA, 323(9): 844–853. The JAMA evaluation behind the $985M median / $1.3B mean figures cited in the funnel-economics section.
-
RAND Corporation (2025). Typical Cost of Developing a New Drug Is Skewed by Few High-Cost Outliers. RAND press release. The 2025 analysis that put the median direct R&D cost at $150M (rising to $708M when capitalized for failures), and showed how a small number of outliers skew the industry-wide mean.
Enjoy Reading This Article?
Here are some more articles you might like to read next:
