How to tell a drug target matters: evidence frameworks for target–disease linkage
A drug candidate can hit its intended protein with favorable potency, selectivity, and pharmacokinetics and still fail in Phase II because the target was never causally linked to the disease in humans. Nelson et al. (2015, Nature Genetics), followed by a revised analysis from King, Davis, and Degner (2019, PLOS Genetics), established that drug programs backed by human genetic evidence are roughly twice as likely to succeed in Phase II and Phase III as programs without it. BIO/Informa industry data over 2011–2020 put biomarker-stratified programs - programs that enroll or analyze patient subgroups defined by a measurable biomarker - at roughly 2× the likelihood of approval versus unstratified ones, a finding that has replicated in subsequent analyses. So evidence for why this target, for this disease, in this patient population carries much of the Phase II burden long before the drug reaches humans.
Concept Translation: Think of a drug target as a feature in a model that you’ve decided to intervene on. Picking a feature that strongly correlates with disease tells you nothing about whether modulating it will change the outcome - that’s the difference between a predictive feature and a causal one. Most of this post is about the evidence techniques the field uses to tell these apart for biological “features” (genes and proteins).
Why target–disease evidence matters in 2026
Industry-wide Phase I likelihood of approval has fallen from about 10.4% to 6.7%. The drivers include a shift toward first-in-class and riskier targets, greater use of biomarkers as surrogate efficacy endpoints (early readouts used as stand-ins for clinical benefit), and tougher regulatory scrutiny. As easy targets are mostly drugged, what remains rests on thinner evidence bases. Biomarker-stratified programs run at roughly double the likelihood of approval of unstratified ones, and targets backed by human genetic evidence run at roughly double the success rate of targets without that support.
Two industry-side facts motivate greater evidence rigor. First, the pipeline is crowded around a small number of targets. About 25% of the global R&D pipeline (roughly 13,600 drug-target pairs) concentrates on just 38 unique biological targets, and in oncology specifically the number of assets per target has grown from 1.8 to 9 over the last two decades (Plenge, 2026). When nine companies are pursuing the same protein, evidence becomes the differentiator - which program has the clearest human-genetic case linking target to disease, the most defensible biomarker, and the cleanest safety picture after accounting for off-target effects (unwanted activity at proteins other than the intended target). Second, human genetic evidence now accompanies approximately two-thirds of recently FDA-approved therapeutics (Ochoa et al., 2022, Nat Rev Drug Discov; Trajanoska et al., 2023, Nature). What was a competitive advantage a decade ago is now table stakes for serious target programs.
What target–disease association evidence actually means
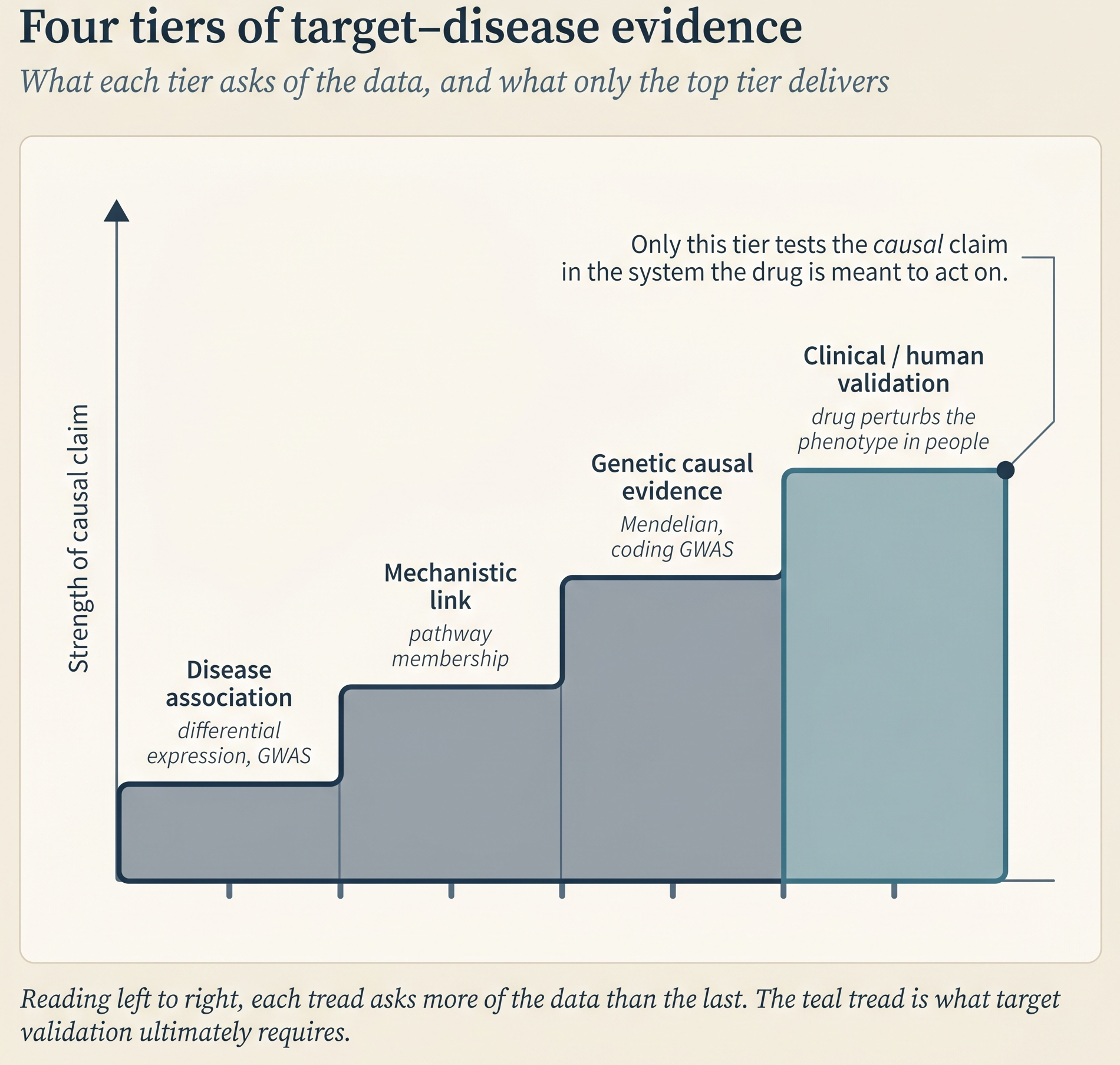
Target-disease evidence is a structured argument built from heterogeneous data sources. A credible argument has four parts:
- A mechanistic hypothesis. A story about how modulating this protein would alter the disease phenotype. We seek a plausible causal chain, going beyond naive correlation with the disease state.
- Human evidence that the hypothesis is correct. Genetic when possible, since human genetic variation imitates the design of a randomized controlled trial without requiring a drug intervention. Multi-omics, patient-tissue, and literature evidence all count, with known caveats.
- Experimental evidence that the modulation produces the therapeutic effect. Knockout, knockdown, or pharmacological perturbation - that is, genetic removal, genetic reduction, or chemical modulation of the target - in disease-relevant models, ideally more than one, that reverses the disease phenotype rather than only altering a biomarker.
- A defensible position on the competing hypothesis. Why this mechanism rather than the three adjacent ones the literature also supports?
Concept Translation: Knockouts and knockdowns are the biology field’s version of ablation studies. Knockout = the gene is removed entirely (full ablation). Knockdown = expression is reduced but not eliminated (partial ablation, more like dropout). Pharmacological perturbation = a chemical inhibits the protein, which is closer to “intervening at inference time” than “retraining without the feature.” All three answer the same question: what does the system do when this component is missing or weakened?
Teams that skip one of these four risk failure. The most common failure is #4 - the team has strong data on its favored mechanism and has never seriously engaged with the alternatives.
Two frameworks for organizing this argument are the AstraZeneca 5R framework and the GOT-IT recommendations.
The AstraZeneca 5R framework: “right target” as the first R

The 5R framework defines five criteria a drug program has to satisfy: right target, right tissue, right safety profile, right patient, right commercial potential. The “right target” criterion is defined narrowly as a strong link between target and disease, predictive biomarkers that identify likely responders, and demonstrated differentiated efficacy relative to alternatives. Lack of efficacy has been the most important cause of project failure in clinical trials. The 5R framework forces teams to separate evidence of target-disease linkage from evidence of tissue availability, safety, and patient stratification (the choice of which patient subgroup to treat). A single piece of data often speaks to only one of these categories, and conflating them makes the evidence base look stronger than it is.
The framework originated at AstraZeneca in the mid-2010s with Cook et al. (2014) Nature Reviews Drug Discovery, followed by Morgan et al. (2018) in Drug Discovery Today, which documented an approximately five-fold improvement in the company’s small-molecule project survival from candidate-selection to Phase III decision after the framework was institutionalized.
GOT-IT: four assessment blocks
The GOT-IT (Good Target Identification and Validation) recommendations, published by Emmerich et al. (2021) in Nature Reviews Drug Discovery, organize target assessment into four blocks. The first is the focus of this post:
- Disease linkage: evaluating the relationship between the target and the disease of interest and determining whether the target is involved in the underlying disease biology.
- Target-related safety: covered in a later post on tissue-specificity.
- Strategic issues: commercial potential, portfolio fit, competition.
- Technical feasibility: covered in a later post on druggability.
5R and GOT-IT overlap, but they aren’t redundant. 5R is a pipeline-level framework, like a checklist for whether a program should advance. GOT-IT is a target-level framework, a checklist for whether the target-assessment work has been done thoroughly. Many organizations use both, sometimes customizing one internally and treating the other as the external benchmark, or using them in tandem with additional frameworks.
Four important distinctions within target–disease linkage
Driver vs passenger mutations
In cancer genomics, driver mutations provide selective advantages for cancer initiation and progression. Passenger mutations accumulate alongside them without contributing to disease fitness. A single gene can host both classes. The APC gene, a highly mutated tumor suppressor in colorectal cancer, carries both driver and passenger mutations across its coding region. Distinguishing them is a computational problem: which variants observed in a tumor caused the malignant phenotype, and which reflect background mutation noise?
Concept Translation: Drivers and passengers are the cancer-genomics version of signal vs. correlated noise. A tumor genome contains hundreds to thousands of mutations, only a handful of which actually move the phenotype. The other mutations co-occur with the drivers because they accumulated in the same lineage. Picking drivers out of the noise is the same kind of feature-selection problem as identifying which input variables actually drive a model’s predictions when the inputs are highly correlated.
A list of “genes mutated in disease X” is raw material. A target-discovery workflow has to rank those genes by probability of being a driver, using statistical tests on mutation frequency against background, functional impact scoring, recurrence across patients, pathway enrichment, and so on. Tools like MutSigCV, OncodriveCLUST, and later deep-learning classifiers exist for this. The output of a cancer-genomics pipeline is a ranked driver list. Moving from that list to a target list requires additional evidence.
A related wrinkle is that some passengers are clinically informative. Weak drivers that fall below detection thresholds, and accumulated “passenger” burden that shifts cell phenotype cumulatively, can matter clinically. Some passengers also provide evolutionary evidence of tumor lineage and can be used for timing inferences.
Causative, supportive, and symptomatic treatment
A second distinction concerns the kind of therapeutic effect a target is meant to produce:
- Causative treatment: therapy directed against the cause of disease. Antiviral agents against SARS-CoV-2. Ritonavir inhibiting HIV protease. A PARP inhibitor in a BRCA-mutated tumor. Targets for causative treatment live upstream in the disease mechanism.
- Symptomatic treatment: therapy that eases symptoms without addressing the underlying cause. Pain relievers. Cough suppressants. Targets for symptomatic treatment live downstream of the disease cause, typically in host response pathways.
- Supportive treatment: care that addresses broader patient needs (physical, existential, palliative) rather than targeting disease biology directly.
For target discovery, this distinction clarifies what “linkage to the disease” has to establish. A target for causative treatment has to be causally upstream of the disease process. A target for symptomatic treatment has to be causally upstream of a symptom, which is a weaker claim. A program’s target-evidence burden depends on which kind of drug it’s trying to develop.
Alzheimer’s provides a standard example of a disease where causative target discovery has resisted easy success. Acetylcholinesterase inhibitors (donepezil, rivastigmine, galantamine) and memantine provide symptomatic therapies that target downstream neurotransmitter biology rather than disease etiology. The amyloid-targeting antibodies aducanumab (accelerated approval 2021, voluntarily withdrawn in 2024), lecanemab (Leqembi, FDA traditional approval July 2023, EMA approval late 2024), and donanemab (Kisunla, FDA approval July 2024) aim at a putative causal mechanism. Phase III data for both lecanemab and donanemab show statistically significant slowing of cognitive decline relative to placebo over 18 months. Debate remains over whether those effect sizes are clinically meaningful. The safety burden of amyloid-related imaging abnormalities (ARIA) has also been a major clinical concern, and the EMA has had a mixed reception of the two drugs. The target-disease linkage evidence for amyloid is therefore stronger than it was a decade ago - there is now Phase III evidence that targeting amyloid produces a measurable clinical effect - but this does not clarify whether amyloid is the causative driver of cognitive decline, a downstream marker, or one of several parallel drivers.1 The earlier generation of amyloid-targeting programs provides a cautionary tale, though continuing studies test whether the hypothesis was right but underpowered, or only partly right with diminishing returns.
Oncogene addiction
Oncogene addiction describes cancer cells’ dependence on individual oncogenes - genes whose altered activity drives tumor growth - to sustain the malignant phenotype. A tumor that depends on an activated oncogene is selectively vulnerable to inhibition of that oncogene. Tumor cells need the oncogenic signal to survive; healthy cells do not. This has been a successful organizing concept in targeted oncology. EGFR mutations in a subset of non-small cell lung cancer, BCR-ABL fusion in chronic myeloid leukemia, and BRAF V600E mutations in melanoma all produce tumors with oncogene-addicted phenotypes, and targeted inhibitors in each context have produced durable clinical responses.
For target-discovery methodology, oncogene addiction is useful because it predicts a specific clinical-trial readout: patients whose tumors harbor the addiction should respond, and others should not. That prediction is testable in Phase II with biomarker-stratified cohorts, and it helps explain why biomarker-stratified oncology programs show higher LOA than unstratified ones (we’ll quantify this in a later post on LOAs by therapeutic area). An evidence package for an oncogene-addiction target should therefore include the candidate biomarker - the measurable feature that marks the likely responder subgroup.
Target-disease correlation vs causal target-disease linkage
The most consequential distinction is also the most abstract. A gene whose expression differs between disease tissue and healthy tissue is associated with the disease. A gene whose loss-of-function variants - variants that reduce or abolish a gene’s activity - segregate with disease risk in human populations at a genetic locus that survives multiple-testing correction, with a plausible protein-level mechanism and Mendelian-randomization-style evidence that inherited variation in the gene affects disease rather than the reverse, is a causal candidate. Curating such evidence is hard. Transcriptomic association is relatively cheap; causal genetic evidence is expensive and sparse.
Concept Translation: Mendelian randomization is the genetics field’s instrumental-variable method. Because alleles are randomly assorted at conception, an inherited variant acts like a randomized treatment assignment - it isn’t influenced by lifestyle, environment, or downstream disease state. If carriers of a loss-of-function variant in gene X are healthier on average, that’s strong evidence that reducing X’s activity reduces disease risk, not just that X is correlated with it. This is closer to an A/B test than to an observational dataset, which is why genetic evidence carries more causal weight than transcriptomic evidence.
Referring back to the 2× headline from earlier, the Nelson et al. (2015) result and the King et al. (2019) revised analysis both confirmed the population-average ~2× clinical-success multiplier for genetically supported drug programs. The same general 2× factor also enriches for labeled side effects, giving it predictive value for toxicology programs (Minikel & Nelson, 2024; Carss et al., 2023). The multiplier is not constant across evidence types. King et al. found that when the causal gene is unambiguous - Mendelian traits, single-gene disorders with clear inheritance patterns, or GWAS associations driven by coding variants where the variant-to-gene mapping is direct - the approval probability multiplier rises above 2× and into the 3× range. The 2026 update from Minikel and Nelson refined this, showing that the multiplier scales with confidence in the causal gene assignment and is largely independent of genetic effect size, minor allele frequency, or year of discovery. In practice, a low-effect-size GWAS hit with confident causal-gene mapping is worth more than a high-effect-size hit at a locus where the variant-to-gene call is ambiguous. Trajanoska et al. (2023, Nature) place this in longer historical context: they identified 40 germline genetic observations that translated directly into approved therapies for 36 rare and 4 common conditions, with a median 25-year interval between target discovery and drug approval. The genetic-anchor strategy works, but it compounds slowly.
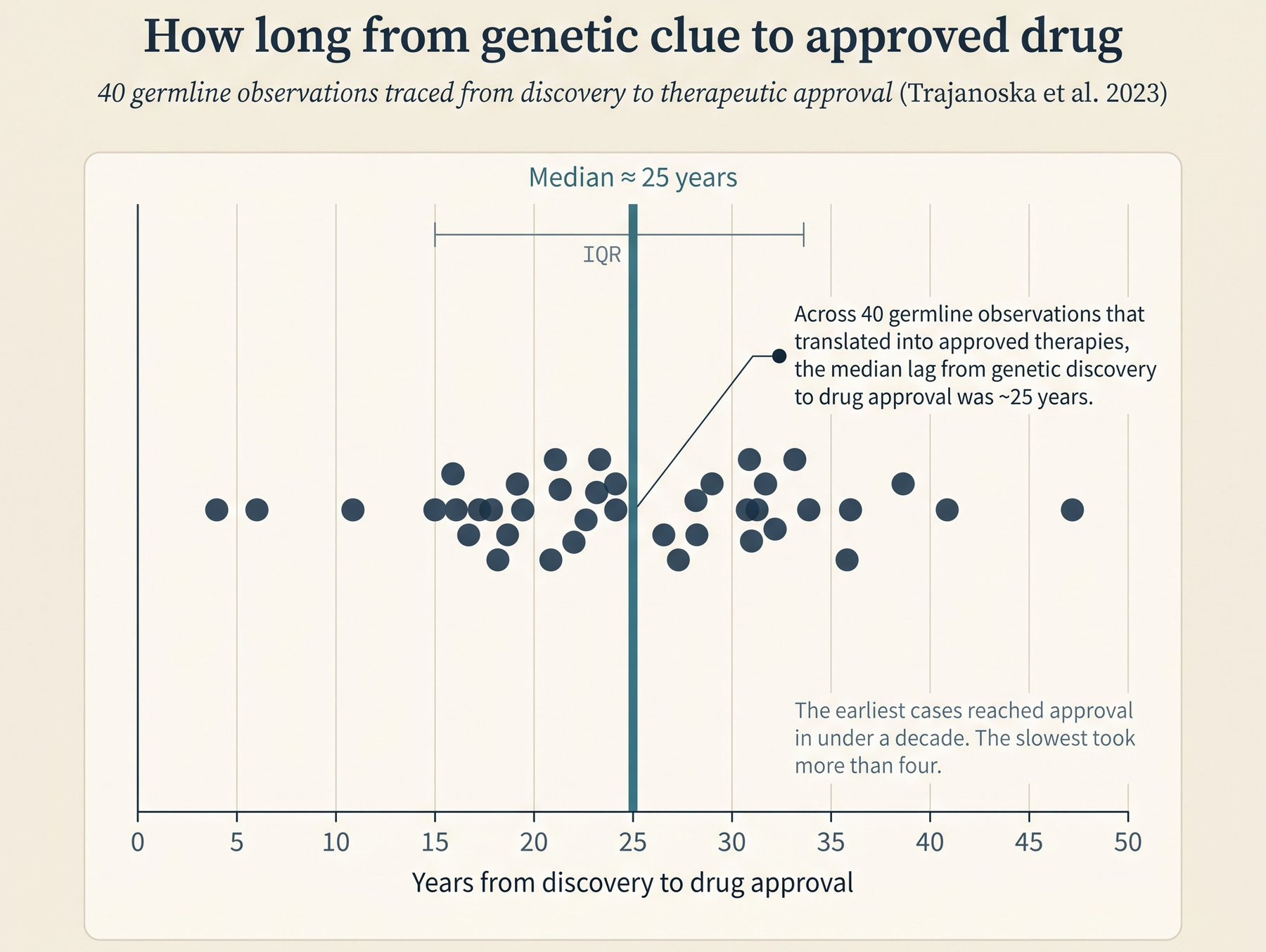
When assembling a target’s evidence package, the quality of the genetic anchor matters more than the quantity of associated data. A coding variant with established protein consequences is worth more than a half-dozen non-coding GWAS signals at adjacent loci with ambiguous fine-mapping.
The omics stack as target–disease evidence

Target-disease linkage evidence comes from an omics stack, and each layer has a characteristic failure mode. Knowing when each layer can mislead you is the core methodological skill.
Genomics and GWAS
Genomics is usually the strongest layer for target-disease linkage. Human genetic variation provides a natural experiment: people who carry a loss-of-function variant in gene X are, on average, similar to people who would be pharmacologically treated with an inhibitor of gene X. If carriers of the variant have lower disease risk, that is a strong argument that inhibiting gene X would lower disease risk too. This is the Mendelian-randomization logic that gives genetic evidence special weight.
Genome-wide association studies (GWAS) are the dominant method for finding these signals. GWAS identifies statistical associations between genetic variants and traits or diseases across large population cohorts. The method has identified thousands of risk loci across hundreds of diseases, now aggregated in resources like the NHGRI-EBI GWAS Catalog.
Concept Translation: GWAS is in some ways a giant feature-importance scan over the genome - for each of millions of variants, test whether allele frequency differs between cases and controls. The genome-wide significance threshold of p < 5×10⁻⁸ is a Bonferroni-style correction for testing roughly a million effective independent variants. The catch is that statistical association picks the neighborhood the causal variant lives in, not the variant itself. Most GWAS hits sit in non-coding DNA - regulatory regions outside protein-coding sequence - which is why follow-up fine-mapping is needed to narrow the signal. Fine-mapping is a feature-attribution problem for a region of the genome: which variant in the correlated cluster is actually doing the work?
The practical limitations matter. GWAS has to apply heavy multiple-testing correction (the classic p < 5×10⁻⁸ genome-wide significance threshold), which means it misses real signals at smaller effect sizes, especially in underpowered cohorts. Most GWAS hits are in non-coding regions, so the protein-level mechanism is often implicit. Identifying the causal gene from a genome-wide-significant locus often requires additional work - fine-mapping (narrowing the locus to the most plausible causal variants), eQTL analysis (testing whether a variant shifts gene expression), and experimental follow-up.
The populations represented in GWAS are also overwhelmingly European-ancestry, which creates a real equity problem when target discovery built on GWAS evidence is applied clinically to more diverse populations. The June 2021 GWAS Catalog inventory put European-ancestry participants at approximately 86.3% of all GWAS subjects, with East Asian at ~5.9% and African at ~1.1%. By 2023 the GWAS Catalog European share had only modestly declined, to around 86.5%, while the GWAS Diversity Monitor, which tracks the running participant counts more broadly, recorded European representation of ~94.5% of total participants by September 2024 (Mills & Rahal, Nat Genet, 2019; Corpas et al., Cell Genomics, 2025). The two metrics differ because the GWAS Diversity Monitor weights large biobank cohorts heavily, but the conclusion is the same: target discovery built on this evidence base will systematically under-serve non-European populations unless deliberately corrected with fine-mapping in diverse cohorts and downstream validation across ancestries. This is a target-evidence problem as much as a deployment problem. A target whose causal gene is well mapped only in European cohorts may not support a defensible target-disease linkage claim for patients of other ancestries until cross-ancestry replication is done.
Concept Translation: This is the same dataset-bias problem that ML practitioners deal with constantly. A model trained on a non-representative dataset can be highly accurate on the in-distribution group while failing on out-of-distribution groups. The fix is the same in principle: collect more representative data, evaluate per-subgroup, and don’t ship a model whose validation doesn’t cover the deployment population.
Transcriptomics
Transcriptomics measures RNA levels, sitting between genotype and protein as a sensitive readout of cellular state. Bulk and single-cell RNA sequencing, along with older microarray data, make up the largest public omics archives (GEO, ArrayExpress) and provide differential-expression evidence: which genes change in disease tissue versus healthy tissue?
The main failure mode of transcriptomics for target discovery is simple. Many transcriptomic changes are bystanders. A gene that changes expression in disease is not necessarily causing the disease; it may be responding to the disease, or changing in parallel with it because of upstream regulation. A transcriptomic hit generates a hypothesis; it does not establish causal linkage. Evidence becomes much stronger when transcriptomics and genetics agree, and the gene is both differentially expressed and under genetic selection in disease.
Concept Translation: Bystander vs. causal in transcriptomics is the same trap as confounded correlation in observational ML. A feature that strongly correlates with the label can still be downstream of the label rather than upstream - predictive of it without driving it. RNA-level changes pick up both kinds and can’t separate them on their own. This is why GWAS-supported transcriptomic hits are weighted more heavily than transcriptomic hits in isolation.
Proteomics and metabolomics
Proteomics measures protein abundance directly. Since most drugs target proteins, proteomic evidence sits closer to the mechanism-of-action question than transcriptomic evidence. The limitations are technical: mass-spec proteomics is expensive, less sensitive than RNA-seq, and struggles with membrane proteins, which include many of the cell-surface receptor and transporter classes that matter most in drug discovery.
Metabolomics measures small-molecule metabolites. It often points to disease-associated metabolic changes that can be traced back to protein targets - which enzymes are producing or consuming these altered metabolites. Metabolomics is particularly useful for metabolic diseases and for target discovery in contexts where disease biology is about pathway flux more than signaling.
Epigenomics
Epigenomics (DNA methylation, histone modifications, chromatin accessibility) provides a reversible regulatory layer above the genome. Epigenetic enzymes are themselves drug targets (HDAC inhibitors, DNMT inhibitors, both with approved drugs), and epigenetic data can identify non-genetic mechanisms driving disease. The characteristic limitation is interpretability: it is often unclear what functional impact a given epigenetic change has on gene expression in the specific disease context.
Concept Translation: A useful mental model: if the genome is the source code, epigenomics is the runtime configuration. Methylation marks and chromatin states change which parts of the code get executed in which cell types, without altering the code itself. Same source, different behavior - and the configuration is reversible, which is why epigenetic enzymes are drug targets.
Multi-omics integration and its limits
Combining these layers sounds straightforward. In practice, it’s a difficult technical problem. Batch effects across studies, confounding by disease severity or patient demographics, differential dropout in single-cell data, and different noise models across modalities all complicate naive integration. The best-case design, when available, analyzes multi-omics data from the same set of patients rather than integrating across cohorts, so sample-level confounders are controlled. That is rarely possible at GWAS-cohort scales, where multi-omics profiling of every participant is prohibitively expensive. Target-discovery workflows that build evidence from public multi-omics resources need to state these limitations explicitly. Workflows that report “our multi-omics integration identified target X” without addressing them are not making a credible argument.
A useful target-disease evidence package usually includes (a) genetic evidence where it exists, as the anchor, (b) transcriptomic and proteomic evidence in disease tissue that is consistent with the genetic signal, (c) tissue-specificity data (see [tissue-specificity evidence] → Tissue specificity: the safety half of target selection2), and (d) network or pathway context that situates the target in plausible biology. No single layer is enough - the layers have to agree.
Networks and knowledge graphs: evidence beyond single-gene associations
A gene-at-a-time view of the omics stack misses mechanistic context. A gene with moderate genetic support and moderate transcriptomic support may look like a weak target in isolation, yet sit at a critical node in a disease-implicated pathway. In that setting, network position is itself evidence. Target-discovery methodology increasingly builds evidence at the network level as well as the gene level.
The method family is biological graphs and knowledge graphs. Protein-protein interaction networks, gene regulatory networks, gene co-expression networks, metabolic networks, and signaling networks each represent a different axis of biological relationship, and each carries evidence relevant to target linkage. Knowledge graphs integrate these with literature-derived associations, drug-target relationships, disease-gene associations, and related data, producing heterogeneous graphs with multiple types of nodes and relationships that can be queried for evidence patterns around a candidate target.
Concept Translation: A biomedical knowledge graph is a heterogeneous graph in the standard ML sense - multiple node types (genes, proteins, diseases, drugs, pathways) and multiple edge types (binds, expressed-in, treats, associated-with). The same graph-ML toolkit applies: node classification (is this gene a likely target?), link prediction (does this gene-disease edge exist but is undiscovered?), graph neural network embeddings, and metapath-based random walks. The biology adds structure but doesn’t change the modeling problem.
Two network features are especially useful for target-linkage arguments. Hubs are highly connected nodes. They are enriched for essential genes and are often promising targets in diseases driven by loss of function, but they are also more likely to be toxic to modulate pharmacologically. Bridging nodes are high-betweenness nodes - nodes that sit on many shortest paths between modules. They are often more attractive targets because their inhibition tends to be non-lethal while still disrupting specific disease-relevant pathways.
Concept Translation: Hubs are nodes with high degree centrality (think high-PageRank pages); knock them out and a lot of the graph collapses, which is why they’re both attractive and dangerous. Bridging nodes are nodes with high betweenness centrality - they sit on many shortest paths, like a bottleneck router connecting subnets. Inhibiting a bridge can disconnect a specific module without crashing the whole network, which is roughly what a clean drug effect looks like.
A knowledge graph over disease-gene-drug-pathway data also lets teams aggregate and score evidence across heterogeneous sources. The Open Targets Platform is the canonical public example: an open-source platform that integrates evidence from genetics (GWAS, ClinVar, ClinGen), somatic mutation data (Cancer Gene Census, COSMIC), known drug-target relationships, expression data, text mining, and animal models, producing a quantitative association score between each gene and each disease. As of the most recent platform release, the Open Targets evidence layer comprises over 27.8 million timestamped evidence assertions across the genes-and-diseases space.
The platform’s scoring methodology matters in practice. The overall target–disease association score is computed as a harmonic sum of source-weighted evidence scores - a scheme in which each additional piece of similar evidence adds less than the previous one, rather than a raw sum or arithmetic mean. The harmonic-sum construction keeps heavily studied targets from accumulating inflated scores through redundant literature citations. Adding the 50th literature reference does not move the score the way the 1st reference does. Within data types that contain multiple sources, the scaling factor uses an inverse-squares construction so that source contributions saturate gracefully rather than letting any single high-throughput source dominate.
Concept Translation: The harmonic-sum aggregator is a regularizer against redundant evidence. Each additional similar source contributes less than the previous one, much like a saturating activation function. Without it, well-studied targets - those with thousands of papers - would dominate the rankings purely by literature volume, and the score would be a popularity ranking instead of an evidence ranking. This is the same problem that plagues citation-count metrics in academia and is solved with the same general technique: diminishing returns on duplicate signal.
Source weights vary by evidence type. ClinGen clinical-validity curations at the “definitive” tier receive an absolute weight of 1.0, the strongest possible single-source evidence. Somatic mutation evidence from the Cancer Gene Census uses a tiered modifying scheme that adjusts the base score by mutational frequency and disease-specificity context: a mutation observed in only 1 sample in the dataset receives a −0.25 modifier (penalizing weak recurrence), while a mutation that occurs more frequently in a particular disease relative to others receives a +0.25 modifier (rewarding disease-specificity). The platform’s “Associations on the Fly” (AOTF) feature, introduced in recent releases, lets users dynamically reweight the evidence contributions to formulate custom therapeutic hypotheses without forking the underlying scoring schema.
The consortium itself is worth noting for field-guide purposes. Open Targets is a pre-competitive public-private partnership - direct competitors contribute to a shared evidence resource before they diverge into proprietary drug programs. As of 2026 the active partner roster includes EMBL-EBI, the Wellcome Sanger Institute, Genentech (Roche Group), GSK, MSD (Merck & Co.), Pfizer, Sanofi, and Bristol Myers Squibb (which joined in late 2022). Biogen and Takeda were earlier members who exited in 2020. The platform aggregates signals that no single company would assemble alone.
Formalizing evidence aggregation as a graph problem also opens the door to graph machine learning. Target-disease association can be formulated as a link-prediction problem on a heterogeneous biomedical knowledge graph: given a graph where some gene-disease edges are known, predict which missing edges are real. Graph neural networks, network-embedding methods, and random-walk-based algorithms have all been applied to this problem. The [knowledge graphs and the rentosertib case study] → Knowledge graphs, network medicine, and the first end-to-end AI-discovered drug picks up this thread in detail.3
A worked example: senolytic target identification in idiopathic pulmonary fibrosis
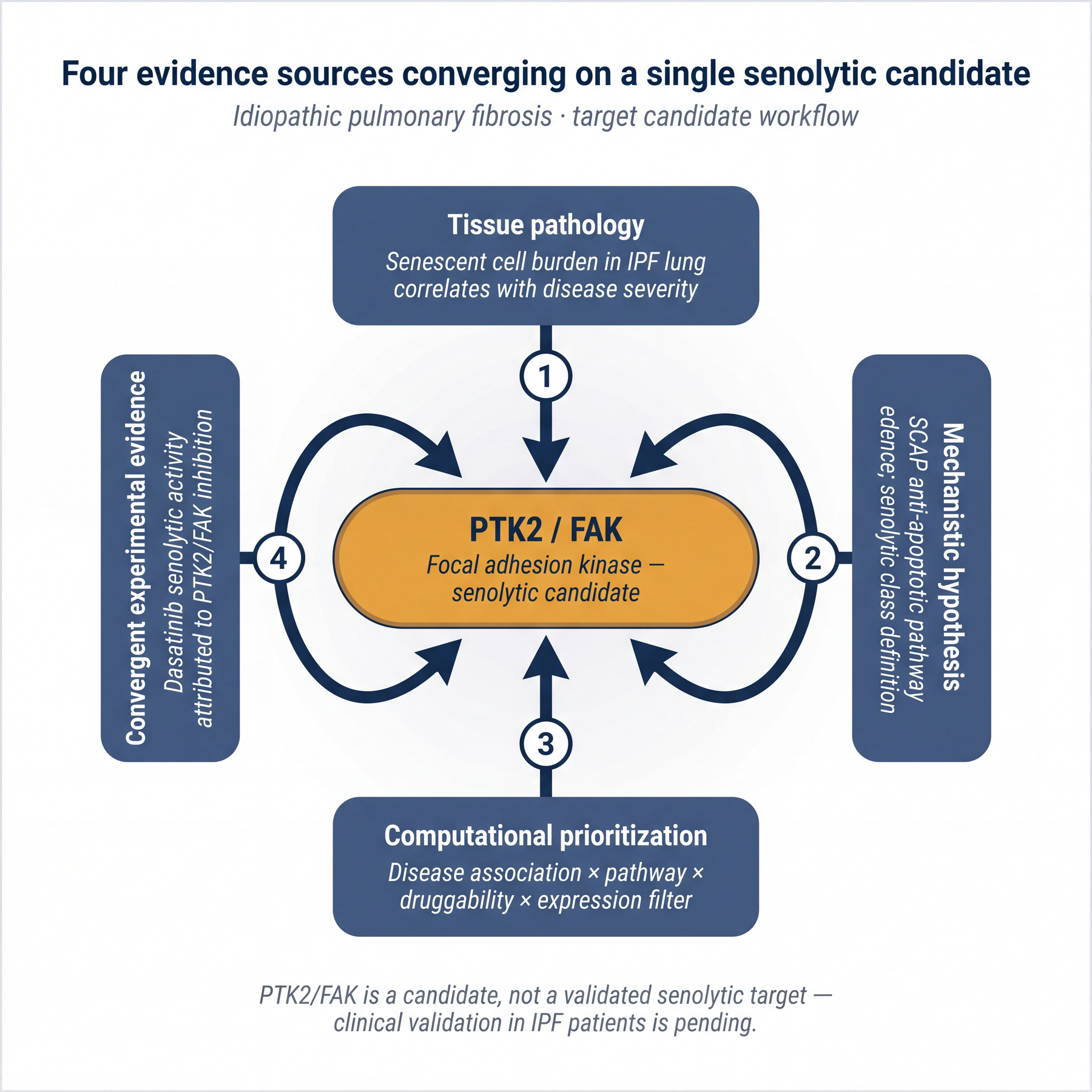
To make the frameworks concrete, consider a target-identification workflow for a specific disease-hypothesis pair: cellular senescence as a target class in idiopathic pulmonary fibrosis (IPF).
The mechanistic hypothesis. Cellular senescence is a permanent state of cell-cycle arrest with a secretory phenotype (SASP) that promotes inflammation and disrupts tissue homeostasis. Senescent cell accumulation is associated with many age-related diseases, and in IPF specifically, senescent cell programs have been documented in disease tissue. The working hypothesis: compounds that selectively kill senescent cells (senolytics) should slow IPF progression by removing the SASP-producing source of inflammation and fibrotic signaling. This is a causative hypothesis in the sense of the earlier framework - senescent cells are positioned as upstream drivers rather than symptoms.
Concept Translation: Senescent cells are sometimes called “zombie cells” - they’ve stopped dividing but won’t die, and they leak inflammatory signals (the SASP) into surrounding tissue. The senolytic strategy is to selectively kill them. From a target-discovery angle, the question is which proteins keep senescent cells alive that are not equally important for keeping healthy cells alive. Those proteins are the senolytic targets.
Evidence layer 1: tissue pathology. Senescent cells are identifiable in IPF lung tissue by senescence-associated markers; their burden correlates with disease severity. This is differential-expression-style evidence - the disease tissue differs from healthy tissue in a specific, characterizable way. It is necessary but insufficient on its own. Senescence could still be a bystander response to lung injury rather than a driver of ongoing fibrosis.
Evidence layer 2: mechanism. Senescent cells are apoptosis-resistant by design; that is part of what makes them persistent. The resistance depends on anti-apoptotic pathways collectively called SCAPs (senescent cell anti-apoptotic pathways). That yields a specific target-class hypothesis: proteins involved in SCAPs are candidate senolytic targets, because inhibiting them should remove the apoptosis resistance and allow senescent cells to die. This narrows the search from “any gene differentially expressed in IPF” to “genes in a specific functional class whose inhibition would have a mechanistically predicted effect.”
Evidence layer 3: computational target prioritization. A knowledge-graph-based target discovery platform can operationalize the combined evidence. The workflow:
- Start from a disease-specific dataset (IPF patient tissue vs healthy controls) and score candidate targets by disease-association strength.
- Apply a gene-set filter restricting candidates to pre-specified functional classes (apoptosis-related pathways, SCAPs).
- Apply a small-molecule druggability filter. This asks whether a target can plausibly be modulated by a conventional small-molecule drug. For senolytic drug development, the delivered modality is small molecule, so candidates that are only tractable by antibody or biologic are de-prioritized.
- Apply an expression filter that focuses on protein classes suitable as drug targets (e.g., kinases, enzymes).
- Apply an optional novelty filter that preferentially surfaces targets with fewer published references, for teams pursuing first-in-class programs against new mechanisms rather than fast-follower strategies against established mechanisms.
The output, checked against known biology. A representative output of this workflow is PTK2 (also known as FAK, focal adhesion kinase) as a high-ranked senolytic candidate. PTK2 is a tyrosine kinase, druggable by small molecules, and expressed in relevant contexts. The biological cross-check comes from independent experimental literature. Dasatinib, a multi-kinase inhibitor originally approved for chronic myeloid leukemia, has been identified as a senolytic in human dermal fibroblasts, and its senolytic activity has been attributed in part to inhibition of PTK2/FAK among other kinases. This independent wet-lab evidence predates the computational prioritization and converges on the same target class.
What this example illustrates. The evidence package for PTK2/FAK as a senolytic target is an aggregation of four sources: disease-tissue pathology (senescent cell burden in IPF), a mechanistic hypothesis (SCAP dependence), computational target prioritization weighted by disease association, pathway membership, and druggability, and convergent experimental evidence from a repurposed clinical-stage compound. No individual layer would be sufficient; together they make a credible argument.
The claim is narrower than it might sound. PTK2 remains a candidate rather than a validated senolytic target in IPF patients; validation would require dosing studies, biomarker responses, and clinical evidence. On the evidence presented here, PTK2 clears the 5R “right target” and GOT-IT “disease linkage” bars for advancing into target validation. Moving from candidate to validated target requires [druggability assessment] → Druggability, ligandability, and modality choice in the AlphaFold 3 era4, and the downstream posts in this series.
Open questions in target–disease evidence work
The benchmarking problem. Target-discovery ML is unusually hard to benchmark. The ground-truth signal - did this target hold up in Phase III? - takes a decade and nine figures per data point. Surrogate benchmarks exist (does the method rediscover known targets for known diseases when given only pre-discovery literature? does it recover genetic signals already in Open Targets? does it predict held-out drug-target relationships?) but all share the same weakness: the evaluation corpus overlaps with the corpus the ML methods were trained on. Truly unbiased benchmarking for target-discovery ML, where the test set is future-clinical rather than past-published, remains a live methodological problem.
The ignorome problem. Research attention across the ~20,000 human protein-coding genes is highly skewed: a small fraction of genes accounts for most of the literature, the annotations, the available assays, and the training data for ML methods. A target-discovery method that only surfaces candidates adjacent to already-well-studied proteins is solving a weaker version of the problem than the one the industry actually needs. The [novelty vs repurposing] → Novelty vs repurposing: when to invent a target and when to reuse one post picks up this thread.5
The causality problem. Most of the evidence stack is associational. Genetic evidence - especially Mendelian-randomization-style evidence - carries more causal weight because it approaches causality in a way other omics layers do not. Methods that can extract more causal structure from non-genetic data (perturbation screens, single-cell perturbation readouts like Perturb-seq, large-scale CRISPR screens) are one of the most active areas of methodological development, and they feed directly into target-disease evidence work. The [synthetic lethality] → Synthetic lethality and combination targets: ML methods for finding drug pairs that work together post gets into CRISPR-screen analysis in depth.6
Evidence frameworks for target-disease linkage separate disciplined target discovery from assertion. Teams that internalize them are less likely to skip steps, and skipped steps are rarely recoverable later. The framework languages (5R, GOT-IT) and the evidence-stack discipline - genomics as anchor, corroborated by transcriptomics and network context, challenged by alternative hypotheses - are what distinguish a defensible target list from one that merely sounds plausible.
Further reading
Foundational framework papers
-
Cook, D., Brown, D., Alexander, R., March, R., Morgan, P., Satterthwaite, G., & Pangalos, M. N. (2014). Lessons learned from the fate of AstraZeneca’s drug pipeline: a five-dimensional framework. Nature Reviews Drug Discovery, 13(6), 419–431. The canonical 5R paper, published from inside AstraZeneca’s R&D leadership at the moment the framework was institutionalized.
-
Morgan, P., Brown, D. G., Lennard, S., Anantharaman, M. J., Gilbert, J., Cook, D., et al. (2018). Impact of a five-dimensional framework on R&D productivity at AstraZeneca. Drug Discovery Today, 23(8), 1395–1399. The quantitative follow-up documenting an approximately five-fold improvement in small-molecule project survival from candidate-selection to Phase III decision after the 5R was institutionalized - the empirical basis for the framework’s enduring methodological status.
-
Emmerich, C. H., Gamboa, L. M., Hofmann, M. C. J., Bonin-Andresen, M., Arbach, O., Schendel, P., et al. (2021). Improving target assessment in biomedical research: the GOT-IT recommendations. Nature Reviews Drug Discovery, 20(1), 64–81. The definitive methodological reference for the four-block target-assessment framework (disease linkage, target-related safety, strategic issues, technical feasibility) that this post’s structure mirrors.
Genetic evidence and the success multiplier
-
Nelson, M. R., Tipney, H., Painter, J. L., Shen, J., Nicoletti, P., Shen, Y., et al. (2015). The support of human genetic evidence for approved drug indications. Nature Genetics, 47(8), 856–860. The original analysis establishing that drug programs with human genetic support are roughly twice as likely to succeed in clinical development. This is the source for the 2× headline that anchors most subsequent literature.
-
King, E. A., Davis, J. W., & Degner, J. F. (2019). Are drug targets with genetic support twice as likely to be approved? Revised estimates of the impact of genetic support for drug mechanisms on the probability of drug approval. PLOS Genetics, 15(12), e1008489. The revised estimate using updated pipeline data, confirming the ~2× general result but showing that high-confidence causal evidence (Mendelian traits, coding-variant GWAS hits) yields a larger >2× multiplier - the source for the 2× / 3× distinction in the post’s hook.
-
Trajanoska, K., Bhérer, C., Taliun, D., Zhou, S., Richards, J. B., & Mooser, V. (2023). From target discovery to clinical drug development with human genetics. Nature, 620(7975), 737–745. Identified 40 germline genetic observations that translated directly into approved therapies for 36 rare and 4 common conditions, with a median 25-year interval between target discovery and drug approval - the long-horizon companion to the multiplier finding.
-
Minikel, E. V., Painter, J. L., Dong, C. C., & Nelson, M. R. (2024). Refining the impact of genetic evidence on clinical success. Nature, 629, 624–629. The 2024 update showing that the genetic-support multiplier scales with confidence in the causal-gene assignment but is largely independent of effect size, allele frequency, or year of discovery.
-
Ochoa, D., Karim, M., Ghoussaini, M., Hulcoop, D. G., McDonagh, E. M., & Dunham, I. (2022). Human genetics evidence supports two-thirds of the 2021 FDA-approved drugs. Nature Reviews Drug Discovery, 21(8), 551. The source for the “two-thirds of recent FDA approvals are underpinned by human genetic evidence” statistic in the introduction.
-
Carss, K. J., Deaton, A. M., Del Rio-Espinola, A., Diogo, D., Fielden, M., Kulkarni, D. A., et al. (2023). Using human genetics to improve safety assessment of therapeutics. Nature Reviews Drug Discovery, 22(2), 145–162. Companion reference for the side-effect-prediction half of the genetic-evidence story.
GWAS diversity
-
Mills, M. C., & Rahal, C. (2019). A scientometric review of genome-wide association studies. Communications Biology, 2, 9. Foundational quantitative analysis of European-ancestry overrepresentation in the GWAS literature; ongoing version available at the GWAS Diversity Monitor (gwasdiversitymonitor.com).
-
Corpas, M., Pius, M., Poburennaya, M., Guio, H., Dwek, M., Nagaraj, S., et al. (2025). Bridging genomics’ greatest challenge: the diversity gap. Cell Genomics, 5(1), 100724. The most recent quantitative update on GWAS diversity, with the September 2024 GWAS Diversity Monitor numbers (~94.5% European participants) and the 2023 GWAS Catalog comparison (~86.5%) used in this post.
Open Targets
-
Open Targets Platform Documentation: target–disease associations and target–disease evidence. platform-docs.opentargets.org. The canonical reference for the harmonic-sum scoring construction, the source-weight schema (including the ClinGen 1.0 absolute weight and the Cancer Gene Census ±0.25 modifiers), and the Associations on the Fly (AOTF) feature. The release-notes page is the right link because evidence-category weighting changes across releases - cite release notes rather than a static platform URL so readers land on the version-of-record for whatever’s current at your publish date.
-
Open Targets news and partner roster: opentargets.org/partners and opentargets.org/news. For the partner-consortium roster cited in the body (EMBL-EBI, Wellcome Sanger, Genentech, GSK, MSD, Pfizer, Sanofi, BMS) and the historical exits (Biogen and Takeda, both 2020).
Anti-amyloid antibodies in Alzheimer’s (for the causative-vs-symptomatic example)
-
van Dyck, C. H., Swanson, C. J., Aisen, P., Bateman, R. J., Chen, C., Gee, M., et al. (2023). Lecanemab in early Alzheimer’s disease. New England Journal of Medicine, 388(1), 9–21. The pivotal Phase III result for lecanemab; the source for the “statistically significant slowing of cognitive decline” claim with the precise CDR-SB effect size.
-
Sims, J. R., Zimmer, J. A., Evans, C. D., Lu, M., Ardayfio, P., Sparks, J., et al. (2023). Donanemab in early symptomatic Alzheimer disease: the TRAILBLAZER-ALZ 2 randomized clinical trial. JAMA, 330(6), 512–527. The pivotal Phase III result for donanemab. The post deliberately frames both drugs as test cases of the amyloid hypothesis rather than as resolved validation of it.
-
For readers who want to follow up, recent controversies around the amyloid hypothesis and the fallout for Alzheimer’s research as a whole are worth a separate dive. We don’t cover the controversy or its causes here. ↩
-
Coming in the next post - how GTEx, Human Protein Atlas, and scRNA-seq sharpen target safety decisions. ↩
-
Coming in a later post - how knowledge graphs get used in end-to-end target-discovery case studies. ↩
-
Coming in the next post - how target candidates get screened for tractability and modality fit. ↩
-
Coming in a later post - how target novelty gets quantified without over-penalizing under-studied biology. ↩
-
Coming in a later post - how CRISPR-screen evidence extends from single targets to target pairs. ↩
Enjoy Reading This Article?
Here are some more articles you might like to read next:
